

 Search and Browse
Search and Browse
 Download
Download
 Enter Data
Enter Data
| Reaction Details | |||
|---|---|---|---|
 | Report a problem with these data | ||
| Target | Insulin-like growth factor 1 receptor | ||
| Ligand | BDBM50158493 | ||
| Substrate/Competitor | n/a | ||
| Meas. Tech. | ChEMBL_1570070 (CHEMBL3789672) | ||
| IC50 | 4.3±n/a nM | ||
| Citation |  Fairhurst, RA; Marsilje, TH; Stutz, S; Boos, A; Niklaus, M; Chen, B; Jiang, S; Lu, W; Furet, P; McCarthy, C; Stauffer, F; Guagnano, V; Vaupel, A; Michellys, PY; Schnell, C; Jeay, S Optimisation of a 5-[3-phenyl-(2-cyclic-ether)-methyl-ether]-4-aminopyrrolopyrimidine series of IGF-1R inhibitors. Bioorg Med Chem Lett26:2057-64 (2016) [PubMed] Article Fairhurst, RA; Marsilje, TH; Stutz, S; Boos, A; Niklaus, M; Chen, B; Jiang, S; Lu, W; Furet, P; McCarthy, C; Stauffer, F; Guagnano, V; Vaupel, A; Michellys, PY; Schnell, C; Jeay, S Optimisation of a 5-[3-phenyl-(2-cyclic-ether)-methyl-ether]-4-aminopyrrolopyrimidine series of IGF-1R inhibitors. Bioorg Med Chem Lett26:2057-64 (2016) [PubMed] Article | ||
| More Info.: | Get all data from this article, Assay Method | ||
| Insulin-like growth factor 1 receptor | |||
| Name: | Insulin-like growth factor 1 receptor | ||
| Synonyms: | CD_antigen=CD221 | IGF-I receptor | IGF1R | IGF1R_HUMAN | Insulin-like growth factor 1 receptor (IGF1R) | Insulin-like growth factor 1 receptor (IGFIR) | Insulin-like growth factor 1 receptor alpha chain | Insulin-like growth factor 1 receptor beta chain | Insulin-like growth factor I receptor | Insulin-like growth factor receptor (IGFR) | ||
| Type: | Protein | ||
| Mol. Mass.: | 154776.79 | ||
| Organism: | Homo sapiens (Human) | ||
| Description: | P08069 | ||
| Residue: | 1367 | ||
| Sequence: |
| ||
| BDBM50158493 | |||
| n/a | |||
| Name | BDBM50158493 | ||
| Synonyms: | CHEMBL3787360 | ||
| Type | Small organic molecule | ||
| Emp. Form. | C28H34FN5O3S | ||
| Mol. Mass. | 539.665 | ||
| SMILES | Nc1ncnc2n(cc(-c3cccc(OCC45CCC(CC4)O5)c3F)c12)[C@@H]1C[C@H](CN2CC[S+]([O-])CC2)C1 |r,wD:26.30,28.33,(-1.03,2.79,;-1.03,1.55,;-2.38,.77,;-2.38,-.77,;-1.03,-1.55,;.3,-.77,;1.76,-1.24,;2.66,.02,;1.76,1.24,;2.23,2.7,;3.76,2.91,;4.33,4.34,;3.38,5.56,;1.86,5.34,;.91,6.55,;1.49,7.98,;.54,9.2,;-.78,8.6,;-1.87,9.66,;-1.5,11.15,;-.02,11.55,;1.07,10.49,;-.08,10.33,;1.29,3.91,;.07,3.74,;.3,.77,;2.24,-2.7,;3.54,-3.45,;2.74,-4.76,;3.1,-6.25,;4.58,-6.69,;4.94,-8.18,;6.42,-8.61,;7.53,-7.55,;8.72,-7.9,;7.17,-6.05,;5.69,-5.62,;1.42,-3.96,)| | ||
| Structure | 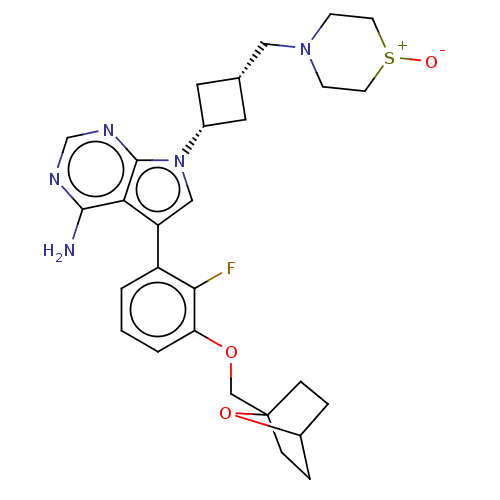
| ||