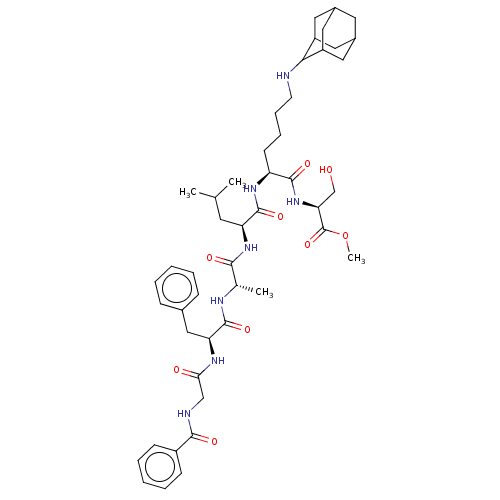
| SMILES | COC(=O)[C@H](CO)NC(=O)[C@H](CCCCNC1C2CC3CC(C2)CC1C3)NC(=O)[C@H](CC(C)C)NC(=O)[C@H](C)NC(=O)[C@H](Cc1ccccc1)NC(=O)CNC(=O)c1ccccc1 |r,wU:42.45,29.32,4.4,wD:37.40,10.10,TLB:15:16:18:22.20.21,THB:20:19:16:22.21.23,20:21:18.19.25:16,23:21:18:25.24.16,23:24:18:22.20.21,(31.33,-35.94,;29.99,-35.18,;28.66,-35.95,;28.66,-37.5,;27.32,-35.19,;27.32,-33.65,;28.65,-32.87,;25.99,-35.96,;24.66,-35.2,;24.65,-33.66,;23.32,-35.97,;23.33,-37.51,;24.67,-38.28,;24.67,-39.82,;26.01,-40.59,;26.01,-42.12,;27.35,-42.89,;27.36,-44.37,;26.16,-45.65,;27.66,-45.23,;29.07,-45.8,;30.08,-44.52,;28.69,-44.86,;30.09,-42.99,;28.7,-42.41,;27.66,-43.65,;21.99,-35.21,;20.66,-35.99,;20.66,-37.53,;19.32,-35.22,;19.31,-33.68,;20.65,-32.91,;20.64,-31.37,;21.98,-33.67,;17.99,-36,;16.65,-35.23,;16.65,-33.69,;15.32,-36.01,;15.33,-37.55,;13.99,-35.24,;12.66,-36.02,;12.66,-37.56,;11.32,-35.25,;11.31,-33.71,;10.55,-32.36,;11.35,-31.04,;10.59,-29.7,;9.05,-29.69,;8.27,-31.02,;9.03,-32.36,;9.99,-36.03,;8.65,-35.26,;8.65,-33.72,;7.32,-36.04,;5.98,-35.27,;4.65,-36.05,;4.66,-37.59,;3.32,-35.28,;3.32,-33.74,;1.98,-32.97,;.65,-33.75,;.66,-35.3,;2,-36.05,)| |
|---|


 Search and Browse
Search and Browse
 Download
Download
 Enter Data
Enter Data

 Stuckey, JI; Simpson, C; Norris-Drouin, JL; Cholensky, SH; Lee, J; Pasca, R; Cheng, N; Dickson, BM; Pearce, KH; Frye, SV; James, LI Structure-Activity Relationships and Kinetic Studies of Peptidic Antagonists of CBX Chromodomains. J Med Chem59:8913-8923 (2016) [PubMed] Article
Stuckey, JI; Simpson, C; Norris-Drouin, JL; Cholensky, SH; Lee, J; Pasca, R; Cheng, N; Dickson, BM; Pearce, KH; Frye, SV; James, LI Structure-Activity Relationships and Kinetic Studies of Peptidic Antagonists of CBX Chromodomains. J Med Chem59:8913-8923 (2016) [PubMed] Article