| Reaction Details |
|---|
 | Report a problem with these data |
| Target | Voltage-dependent L-type calcium channel subunit alpha-1S |
|---|
| Ligand | BDBM50226128 |
|---|
| Substrate/Competitor | n/a |
|---|
| Meas. Tech. | ChEMBL_42923 (CHEMBL654566) |
|---|
| Ki | 0.205000±n/a nM |
|---|
| Citation |  Tamazawa, K; Arima, H; Kojima, T; Isomura, Y; Okada, M; Fujita, S; Furuya, T; Takenaka, T; Inagaki, O; Terai, M Stereoselectivity of a potent calcium antagonist, 1-benzyl-3-pyrrolidinyl methyl 2,6-dimethyl-4-(m-nitrophenyl)-1,4-dihydropyridine-3,5-dicarboxylate. J Med Chem29:2504-11 (1986) [PubMed] Tamazawa, K; Arima, H; Kojima, T; Isomura, Y; Okada, M; Fujita, S; Furuya, T; Takenaka, T; Inagaki, O; Terai, M Stereoselectivity of a potent calcium antagonist, 1-benzyl-3-pyrrolidinyl methyl 2,6-dimethyl-4-(m-nitrophenyl)-1,4-dihydropyridine-3,5-dicarboxylate. J Med Chem29:2504-11 (1986) [PubMed] |
|---|
| More Info.: | Get all data from this article, Assay Method |
|---|
| |
| Voltage-dependent L-type calcium channel subunit alpha-1S |
|---|
| Name: | Voltage-dependent L-type calcium channel subunit alpha-1S |
|---|
| Synonyms: | CAC1S_RAT | Cach1 | Cacn1 | Cacna1s | Cacnl1a3 | Calcium channel, L type, alpha-1 polypeptide, isoform 3, skeletal muscle | Cchl1a3 | ROB1 | Voltage-gated L-type calcium channel | Voltage-gated L-type calcium channel alpha-1S subunit | Voltage-gated calcium channel subunit alpha Cav1.1 |
|---|
| Type: | PROTEIN |
|---|
| Mol. Mass.: | 210392.58 |
|---|
| Organism: | Rattus norvegicus |
|---|
| Description: | ChEMBL_92777 |
|---|
| Residue: | 1850 |
|---|
| Sequence: | MEPSSPQDEGLRKKQPKKPVPEILPRPPRALFCLTLQNPLRKACISVVEWKPFETIILLT
IFANCVALAVYLPMPEDDNNTLNLGLEKLEYFFLIVFSIEAAMKIIAYGFLFHQDAYLRS
GWNVLDFIIVFLGVFTAILEQVNIIQTNTAPMSSKGAGLDVKALRAFRVLRPLRLVSGVP
SLQVVLNSIFKAMLPLFHIALLVLFMVIIYAIIGLELFKGKMHKTCYFIGTDIVATVENE
KPSPCARTGSGRPCTINGSECRGGWPGPNHGITHFDNFGFSMLTVYQCISMEGWTDVLYW
VNDAIGNEWPWIYFVTLILLGSFFILNLVLGVLSGEFTKEREKAKSRGTFQKLREKQQLE
EDLRGYMSWITQGEVMDVDDLREGKLSLDEGGSDTESLYEIEGLNKIIQFIRHWRQWNRV
FRWKCHDLVKSKVFYWLVILIVALNTLSIASEHHNQPLWLTHLQDVANRVLLALFTIEML
MKMYGLGLRQYFMSIFNRFDCFVVCSGILEILLVESGAMTPLGISVLRCIRLLRLFKITK
YWTSLSNLVASLLNSIRSIASLLLLLFLFMIIFALLGMQLFGGRYDFEDTEVRRSNFDNF
PQALISVFQVLTGEDWNSVMYNGIMAYGGPSYPGVLVCIYFIILFVCGNYILLNVFLAIA
VDNLAEAESLTSAQKAKAEERKRRKMSRGLPDKSEEERSTMTKKLEQKPKGEGIPTTAKL
KIDEFESNVNEVKDPYPSADFPGDDEEDEPEIPASPRPRPLAELQLKEKAVPIPEASSFF
IFSPTNKIRVLCHRIVNATWFTNFILLFILLSSAALAAEDPIRADSMRNQILEYFDYVFT
AVFTVEIVLKMTTYGAFLHKGSFCRNYFNILDLLVVAVSLISMGLESSAISVVKILRVLR
VLRPLRAINRAKGLKHVVQCVFVAIRTIGNIVLVTTLLQFMFACIGVQLFKGKFYSCNDL
SKMTEEECRGYYYIYKDGDPTQIELRPRQWIHNDFHFDNVLSAMMSLFTVSTFEGWPQLL
YKAIDSNEEDTGPVYNNRVEMAIFFIIYIILIAFFMMNIFVGFVIVTFQEQGETEYKNCE
LDKNQRQCVQYALKARPLRCYIPKNPYQYQVWYVVTSSYFEYLMFALIMLNTICLGMQHY
NQSEQMNHISDILNVAFTIIFTLEMILKLIAFKPRGYFGDPWNVFDFLIVIGSIIDVILS
EIDTLLASSGGLYCLGGGCGNVDPDESARISSAFFRLFRVMRLIKLLSRAEGVRTLLWTF
IKSFQALPYVALLIVMLFFIYAVIGMQMFGKIAMVDGTQINRNNNFQTFPQAVLLLFRCA
TGEAWQEILLACSYGKRCDPESDYAPGEEYACGTNFAYYYFISFYMLCAFLIINLFVAVI
MDNFDYLTRDWSILGPHHLDEFKAIWAEYDPEAKGRIKHLDVVTLLRRIQPPLGFGKFCP
HRVACKRLVGMNMPLNSDGTVTFNATLFALVRTALKIKTEGNFEQANEELRAIIKKIWKR
TSMKLLDQVIPPIGDDEVTVGKFYATFLIQEHFRKFMKRQEEYYGYRPKKDTVQIQAGLR
TIEEEAAPEIHRAISGDLTAEEELERAMVEAAMEEGIFRRTGGLFGQVDNFLERTNSLPP
VMANQRPLQFAEMEMEELESPVFLEDFPQNPGTHPLARANTNNANANVAYGNSSHRNSPV
FSSIRYERELLEEAGRPVTREGPFSQPCSVSGVNSRSHVDKLERQMSQRRMPKGQVPPSP
CQLSQEKHPVHEEGKGPRSWSTETSDSESFEERVPRNSAHKCTAPATTMLIQEALVRGGL
DSLAADANFVMATGQALADACQMEPEEVEVAATELLKRESPKGGAMPREP
|
|
|
|---|
| BDBM50226128 |
|---|
| n/a |
|---|
| Name | BDBM50226128 |
|---|
| Synonyms: | CHEMBL2093893 |
|---|
| Type | Small organic molecule |
|---|
| Emp. Form. | C27H30ClN3O6 |
|---|
| Mol. Mass. | 527.997 |
|---|
| SMILES | Cl.[H][C@@]1(CCN(Cc2ccccc2)C1)OC(=O)C1=C(C)N=C(C)\C(=C(\O)OC)[C@]1([H])c1cccc(c1)[N+]([O-])=O |r,c:18,t:21| |
|---|
| Structure | 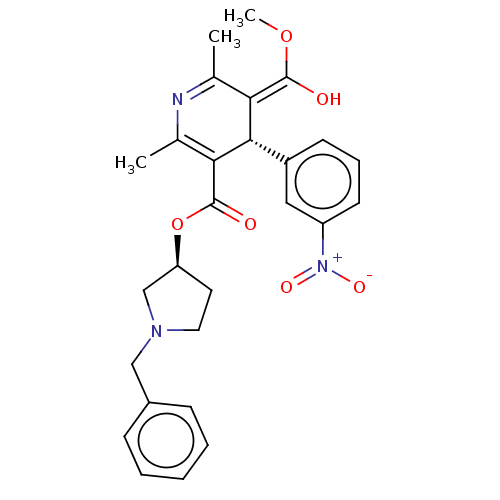
|
|---|


 Search and Browse
Search and Browse
 Download
Download
 Enter Data
Enter Data

 Tamazawa, K; Arima, H; Kojima, T; Isomura, Y; Okada, M; Fujita, S; Furuya, T; Takenaka, T; Inagaki, O; Terai, M Stereoselectivity of a potent calcium antagonist, 1-benzyl-3-pyrrolidinyl methyl 2,6-dimethyl-4-(m-nitrophenyl)-1,4-dihydropyridine-3,5-dicarboxylate. J Med Chem29:2504-11 (1986) [PubMed]
Tamazawa, K; Arima, H; Kojima, T; Isomura, Y; Okada, M; Fujita, S; Furuya, T; Takenaka, T; Inagaki, O; Terai, M Stereoselectivity of a potent calcium antagonist, 1-benzyl-3-pyrrolidinyl methyl 2,6-dimethyl-4-(m-nitrophenyl)-1,4-dihydropyridine-3,5-dicarboxylate. J Med Chem29:2504-11 (1986) [PubMed]