| Reaction Details |
|---|
 | Report a problem with these data |
| Target | Protein mono-ADP-ribosyltransferase PARP4 |
|---|
| Ligand | BDBM50234915 |
|---|
| Substrate/Competitor | n/a |
|---|
| Meas. Tech. | ChEMBL_1654154 (CHEMBL4003520) |
|---|
| IC50 | 64565±n/a nM |
|---|
| Citation |  Nathubhai, A; Haikarainen, T; Koivunen, J; Murthy, S; Koumanov, F; Lloyd, MD; Holman, GD; Pihlajaniemi, T; Tosh, D; Lehti�, L; Threadgill, MD Highly Potent and Isoform Selective Dual Site Binding Tankyrase/Wnt Signaling Inhibitors That Increase Cellular Glucose Uptake and Have Antiproliferative Activity. J Med Chem60:814-820 (2017) [PubMed] Article Nathubhai, A; Haikarainen, T; Koivunen, J; Murthy, S; Koumanov, F; Lloyd, MD; Holman, GD; Pihlajaniemi, T; Tosh, D; Lehti�, L; Threadgill, MD Highly Potent and Isoform Selective Dual Site Binding Tankyrase/Wnt Signaling Inhibitors That Increase Cellular Glucose Uptake and Have Antiproliferative Activity. J Med Chem60:814-820 (2017) [PubMed] Article |
|---|
| More Info.: | Get all data from this article, Assay Method |
|---|
| |
| Protein mono-ADP-ribosyltransferase PARP4 |
|---|
| Name: | Protein mono-ADP-ribosyltransferase PARP4 |
|---|
| Synonyms: | (ARTD4 or PARP4) | 193 kDa vault protein | 2.4.2.- | 2.4.2.30 | ADP-ribosyltransferase diphtheria toxin-like 4 | ADPRTL1 | ADPRTL1 | ARTD4 | KIAA0177 | KIAA0177 GN | PARP-4 | PARP-related/IalphaI-related H5/proline-rich | PARP4 | PARP4_HUMAN | PARPL | PH5P | Poly [ADP-ribose] polymerase 4 | Synonyms=ADPRTL1 | VPARP | Vault poly(ADP-ribose) polymerase |
|---|
| Type: | n/a |
|---|
| Mol. Mass.: | 192563.23 |
|---|
| Organism: | Homo sapiens (Human) |
|---|
| Description: | Q9UKK3 |
|---|
| Residue: | 1724 |
|---|
| Sequence: | MVMGIFANCIFCLKVKYLPQQQKKKLQTDIKENGGKFSFSLNPQCTHIILDNADVLSQYQ
LNSIQKNHVHIANPDFIWKSIREKRLLDVKNYDPYKPLDITPPPDQKASSSEVKTEGLCP
DSATEEEDTVELTEFGMQNVEIPHLPQDFEVAKYNTLEKVGMEGGQEAVVVELQCSRDSR
DCPFLISSHFLLDDGMETRRQFAIKKTSEDASEYFENYIEELKKQGFLLREHFTPEATQL
ASEQLQALLLEEVMNSSTLSQEVSDLVEMIWAEALGHLEHMLLKPVNRISLNDVSKAEGI
LLLVKAALKNGETAEQLQKMMTEFYRLIPHKGTMPKEVNLGLLAKKADLCQLIRDMVNVC
ETNLSKPNPPSLAKYRALRCKIEHVEQNTEEFLRVRKEVLQNHHSKSPVDVLQIFRVGRV
NETTEFLSKLGNVRPLLHGSPVQNIVGILCRGLLLPKVVEDRGVQRTDVGNLGSGIYFSD
SLSTSIKYSHPGETDGTRLLLICDVALGKCMDLHEKDFSLTEAPPGYDSVHGVSQTASVT
TDFEDDEFVVYKTNQVKMKYIIKFSMPGDQIKDFHPSDHTELEEYRPEFSNFSKVEDYQL
PDAKTSSSTKAGLQDASGNLVPLEDVHIKGRIIDTVAQVIVFQTYTNKSHVPIEAKYIFP
LDDKAAVCGFEAFINGKHIVGEIKEKEEAQQEYLEAVTQGHGAYLMSQDAPDVFTVSVGN
LPPKAKVLIKITYITELSILGTVGVFFMPATVAPWQQDKALNENLQDTVEKICIKEIGTK
QSFSLTMSIEMPYVIEFIFSDTHELKQKRTDCKAVISTMEGSSLDSSGFSLHIGLSAAYL
PRMWVEKHPEKESEACMLVFQPDLDVDLPDLASESEVIICLDCSSSMEGVTFLQAKQIAL
HALSLVGEKQKVNIIQFGTGYKELFSYPKHITSNTMAAEFIMSATPTMGNTDFWKTLRYL
SLLYPARGSRNILLVSDGHLQDESLTLQLVKRSRPHTRLFACGIGSTANRHVLRILSQCG
AGVFEYFNAKSKHSWRKQIEDQMTRLCSPSCHSVSVKWQQLNPDVPEALQAPAQVPSLFL
NDRLLVYGFIPHCTQATLCALIQEKEFRTMVSTTELQKTTGTMIHKLAARALIRDYEDGI
LHENETSHEMKKQTLKSLIIKLSKENSLITQFTSFVAVEKRDENESPFPDIPKVSELIAK
EDVDFLPYMSWQGEPQEAVRNQSLLASSEWPELRLSKRKHRKIPFSKRKMELSQPEVSED
FEEDGLGVLPAFTSNLERGGVEKLLDLSWTESCKPTATEPLFKKVSPWETSTSSFFPILA
PAVGSYLPPTARAHSPASLSFASYRQVASFGSAAPPRQFDASQFSQGPVPGTCADWIPQS
ASCPTGPPQNPPSSPYCGIVFSGSSLSSAQSAPLQHPGGFTTRPSAGTFPELDSPQLHFS
LPTDPDPIRGFGSYHPSASSPFHFQPSAASLTANLRLPMASALPEALCSQSRTTPVDLCL
LEESVGSLEGSRCPVFAFQSSDTESDELSEVLQDSCFLQIKCDTKDDSILCFLEVKEEDE
IVCIQHWQDAVPWTELLSLQTEDGFWKLTPELGLILNLNTNGLHSFLKQKGIQSLGVKGR
ECLLDLIATMLVLQFIRTRLEKEGIVFKSLMKMDDASISRNIPWAFEAIKQASEWVRRTE
GQYPSICPRLELGNDWDSATKQLLGLQPISTVSPLHRVLHYSQG
|
|
|
|---|
| BDBM50234915 |
|---|
| n/a |
|---|
| Name | BDBM50234915 |
|---|
| Synonyms: | CHEMBL4069447 |
|---|
| Type | Small organic molecule |
|---|
| Emp. Form. | C28H23N5O3 |
|---|
| Mol. Mass. | 477.5139 |
|---|
| SMILES | Cc1cccc2c1nc(CCC(=O)Nc1ccc(cc1)C(=O)Nc1cccc3cccnc13)[nH]c2=O |
|---|
| Structure | 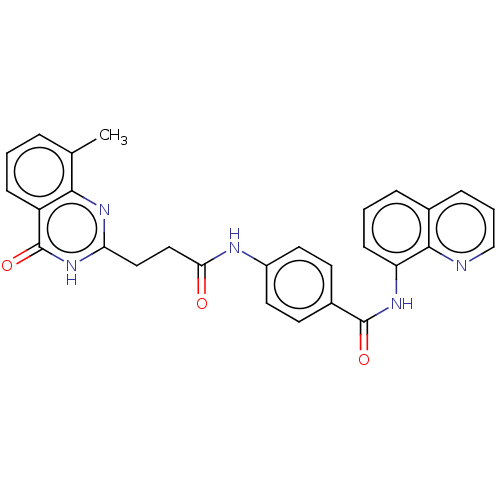
|
|---|


 Search and Browse
Search and Browse
 Download
Download
 Enter Data
Enter Data

 Nathubhai, A; Haikarainen, T; Koivunen, J; Murthy, S; Koumanov, F; Lloyd, MD; Holman, GD; Pihlajaniemi, T; Tosh, D; Lehti�, L; Threadgill, MD Highly Potent and Isoform Selective Dual Site Binding Tankyrase/Wnt Signaling Inhibitors That Increase Cellular Glucose Uptake and Have Antiproliferative Activity. J Med Chem60:814-820 (2017) [PubMed] Article
Nathubhai, A; Haikarainen, T; Koivunen, J; Murthy, S; Koumanov, F; Lloyd, MD; Holman, GD; Pihlajaniemi, T; Tosh, D; Lehti�, L; Threadgill, MD Highly Potent and Isoform Selective Dual Site Binding Tankyrase/Wnt Signaling Inhibitors That Increase Cellular Glucose Uptake and Have Antiproliferative Activity. J Med Chem60:814-820 (2017) [PubMed] Article