| Reaction Details |
|---|
 | Report a problem with these data |
| Target | Prostacyclin receptor |
|---|
| Ligand | BDBM50235383 |
|---|
| Substrate/Competitor | n/a |
|---|
| Meas. Tech. | ChEMBL_1655797 (CHEMBL4005267) |
|---|
| IC50 | 62±n/a nM |
|---|
| Citation |  Tran, TA; Kramer, B; Shin, YJ; Vallar, P; Boatman, PD; Zou, N; Sage, CR; Gharbaoui, T; Krishnan, A; Pal, B; Shakya, SR; Garrido Montalban, A; Adams, JW; Ramirez, J; Behan, DP; Shifrina, A; Blackburn, A; Leakakos, T; Shi, Y; Morgan, M; Sadeque, A; Chen, W; Unett, DJ; Gaidarov, I; Chen, X; Chang, S; Shu, HH; Tung, SF; Semple, G Discovery of 2-(((1r,4r)-4-(((4-Chlorophenyl)(phenyl)carbamoyl)oxy)methyl)cyclohexyl)methoxy)acetate (Ralinepag): An Orally Active Prostacyclin Receptor Agonist for the Treatment of Pulmonary Arterial Hypertension. J Med Chem60:913-927 (2017) [PubMed] Article Tran, TA; Kramer, B; Shin, YJ; Vallar, P; Boatman, PD; Zou, N; Sage, CR; Gharbaoui, T; Krishnan, A; Pal, B; Shakya, SR; Garrido Montalban, A; Adams, JW; Ramirez, J; Behan, DP; Shifrina, A; Blackburn, A; Leakakos, T; Shi, Y; Morgan, M; Sadeque, A; Chen, W; Unett, DJ; Gaidarov, I; Chen, X; Chang, S; Shu, HH; Tung, SF; Semple, G Discovery of 2-(((1r,4r)-4-(((4-Chlorophenyl)(phenyl)carbamoyl)oxy)methyl)cyclohexyl)methoxy)acetate (Ralinepag): An Orally Active Prostacyclin Receptor Agonist for the Treatment of Pulmonary Arterial Hypertension. J Med Chem60:913-927 (2017) [PubMed] Article |
|---|
| More Info.: | Get all data from this article, Assay Method |
|---|
| |
| Prostacyclin receptor |
|---|
| Name: | Prostacyclin receptor |
|---|
| Synonyms: | PGI receptor | PI2R_HUMAN | PRIPR | PTGIR | Prostacyclin (IP) Receptor | Prostacyclin receptor | Prostaglandin I | Prostaglandin I2 | Prostaglandin I2 receptor | Prostanoid IP receptor |
|---|
| Type: | G Protein-Coupled Receptor (GPCR) |
|---|
| Mol. Mass.: | 40968.22 |
|---|
| Organism: | Homo sapiens (Human) |
|---|
| Description: | The membranes prepared from human platelet were used in binding assay. |
|---|
| Residue: | 386 |
|---|
| Sequence: | MADSCRNLTYVRGSVGPATSTLMFVAGVVGNGLALGILSARRPARPSAFAVLVTGLAATD
LLGTSFLSPAVFVAYARNSSLLGLARGGPALCDAFAFAMTFFGLASMLILFAMAVERCLA
LSHPYLYAQLDGPRCARLALPAIYAFCVLFCALPLLGLGQHQQYCPGSWCFLRMRWAQPG
GAAFSLAYAGLVALLVAAIFLCNGSVTLSLCRMYRQQKRHQGSLGPRPRTGEDEVDHLIL
LALMTVVMAVCSLPLTIRCFTQAVAPDSSSEMGDLLAFRFYAFNPILDPWVFILFRKAVF
QRLKLWVCCLCLGPAHGDSQTPLSQLASGRRDPRAPSAPVGKEGSCVPLSAWGEGQVEPL
PPTQQSSGSAVGTSSKAEASVACSLC
|
|
|
|---|
| BDBM50235383 |
|---|
| n/a |
|---|
| Name | BDBM50235383 |
|---|
| Synonyms: | ACT-293987 | NS-304 | Selexipag | Uptravi |
|---|
| Type | Small organic molecule |
|---|
| Emp. Form. | C26H32N4O4S |
|---|
| Mol. Mass. | 496.622 |
|---|
| SMILES | CC(C)N(CCCCOCC(=O)NS(C)(=O)=O)c1cnc(-c2ccccc2)c(n1)-c1ccccc1 |
|---|
| Structure | 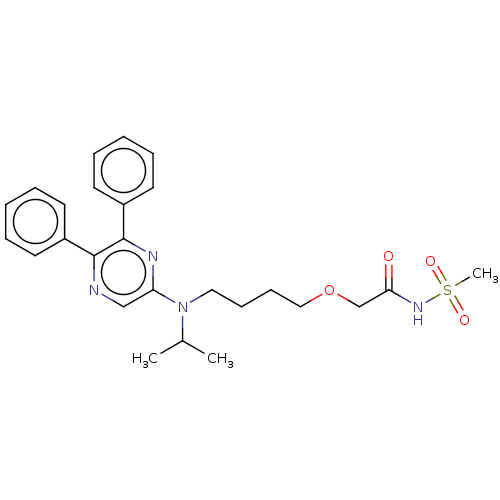
|
|---|


 Search and Browse
Search and Browse
 Download
Download
 Enter Data
Enter Data

 Tran, TA; Kramer, B; Shin, YJ; Vallar, P; Boatman, PD; Zou, N; Sage, CR; Gharbaoui, T; Krishnan, A; Pal, B; Shakya, SR; Garrido Montalban, A; Adams, JW; Ramirez, J; Behan, DP; Shifrina, A; Blackburn, A; Leakakos, T; Shi, Y; Morgan, M; Sadeque, A; Chen, W; Unett, DJ; Gaidarov, I; Chen, X; Chang, S; Shu, HH; Tung, SF; Semple, G Discovery of 2-(((1r,4r)-4-(((4-Chlorophenyl)(phenyl)carbamoyl)oxy)methyl)cyclohexyl)methoxy)acetate (Ralinepag): An Orally Active Prostacyclin Receptor Agonist for the Treatment of Pulmonary Arterial Hypertension. J Med Chem60:913-927 (2017) [PubMed] Article
Tran, TA; Kramer, B; Shin, YJ; Vallar, P; Boatman, PD; Zou, N; Sage, CR; Gharbaoui, T; Krishnan, A; Pal, B; Shakya, SR; Garrido Montalban, A; Adams, JW; Ramirez, J; Behan, DP; Shifrina, A; Blackburn, A; Leakakos, T; Shi, Y; Morgan, M; Sadeque, A; Chen, W; Unett, DJ; Gaidarov, I; Chen, X; Chang, S; Shu, HH; Tung, SF; Semple, G Discovery of 2-(((1r,4r)-4-(((4-Chlorophenyl)(phenyl)carbamoyl)oxy)methyl)cyclohexyl)methoxy)acetate (Ralinepag): An Orally Active Prostacyclin Receptor Agonist for the Treatment of Pulmonary Arterial Hypertension. J Med Chem60:913-927 (2017) [PubMed] Article