| Reaction Details |
|---|
 | Report a problem with these data |
| Target | Metabotropic glutamate receptor 2 |
|---|
| Ligand | BDBM17660 |
|---|
| Substrate/Competitor | n/a |
|---|
| Ki | >10000±n/a nM |
|---|
| Comments | PDSP_1580 |
|---|
| Citation |  Schaffhauser, H; Richards, JG; Cartmell, J; Chaboz, S; Kemp, JA; Klingelschmidt, A; Messer, J; Stadler, H; Woltering, T; Mutel, V In vitro binding characteristics of a new selective group II metabotropic glutamate receptor radioligand, [3H]LY354740, in rat brain. Mol Pharmacol53:228-33 (1998) [PubMed] Schaffhauser, H; Richards, JG; Cartmell, J; Chaboz, S; Kemp, JA; Klingelschmidt, A; Messer, J; Stadler, H; Woltering, T; Mutel, V In vitro binding characteristics of a new selective group II metabotropic glutamate receptor radioligand, [3H]LY354740, in rat brain. Mol Pharmacol53:228-33 (1998) [PubMed] |
|---|
| More Info.: | Get all data from this article |
|---|
| |
| Metabotropic glutamate receptor 2 |
|---|
| Name: | Metabotropic glutamate receptor 2 |
|---|
| Synonyms: | GPRC1B | GRM2 | GRM2_HUMAN | MGLUR2 | Metabotropic glutamate receptor | glutamate receptor, metabotropic 2 precursor | mGlu2 | metabotropic glutamate 2 |
|---|
| Type: | Enzyme |
|---|
| Mol. Mass.: | 95584.88 |
|---|
| Organism: | Homo sapiens (Human) |
|---|
| Description: | Q14416 |
|---|
| Residue: | 872 |
|---|
| Sequence: | MGSLLALLALLLLWGAVAEGPAKKVLTLEGDLVLGGLFPVHQKGGPAEDCGPVNEHRGIQ
RLEAMLFALDRINRDPHLLPGVRLGAHILDSCSKDTHALEQALDFVRASLSRGADGSRHI
CPDGSYATHGDAPTAITGVIGGSYSDVSIQVANLLRLFQIPQISYASTSAKLSDKSRYDY
FARTVPPDFFQAKAMAEILRFFNWTYVSTVASEGDYGETGIEAFELEARARNICVATSEK
VGRAMSRAAFEGVVRALLQKPSARVAVLFTRSEDARELLAASQRLNASFTWVASDGWGAL
ESVVAGSEGAAEGAITIELASYPISDFASYFQSLDPWNNSRNPWFREFWEQRFRCSFRQR
DCAAHSLRAVPFEQESKIMFVVNAVYAMAHALHNMHRALCPNTTRLCDAMRPVNGRRLYK
DFVLNVKFDAPFRPADTHNEVRFDRFGDGIGRYNIFTYLRAGSGRYRYQKVGYWAEGLTL
DTSLIPWASPSAGPLPASRCSEPCLQNEVKSVQPGEVCCWLCIPCQPYEYRLDEFTCADC
GLGYWPNASLTGCFELPQEYIRWGDAWAVGPVTIACLGALATLFVLGVFVRHNATPVVKA
SGRELCYILLGGVFLCYCMTFIFIAKPSTAVCTLRRLGLGTAFSVCYSALLTKTNRIARI
FGGAREGAQRPRFISPASQVAICLALISGQLLIVVAWLVVEAPGTGKETAPERREVVTLR
CNHRDASMLGSLAYNVLLIALCTLYAFKTRKCPENFNEAKFIGFTMYTTCIIWLAFLPIF
YVTSSDYRVQTTTMCVSVSLSGSVVLGCLFAPKLHIILFQPQKNVVSHRAPTSRFGSAAA
RASSSLGQGSGSQFVPTVCNGREVVDSTTSSL
|
|
|
|---|
| BDBM17660 |
|---|
| n/a |
|---|
| Name | BDBM17660 |
|---|
| Synonyms: | (2S)-2-amino-3-(3,5-dioxo-1,2,4-oxadiazolidin-2-yl)propanoic acid | 2-amino-3-(3,5-dioxo[1,2,4]oxadiazolidin-2-yl)propionic acid | CHEMBL279956 | Quisqualate | Quisqualate, L | quisqualic acid |
|---|
| Type | Small organic molecule |
|---|
| Emp. Form. | C5H7N3O5 |
|---|
| Mol. Mass. | 189.1262 |
|---|
| SMILES | N[C@@H](Cn1oc(=O)[nH]c1=O)C(O)=O |
|---|
| Structure | 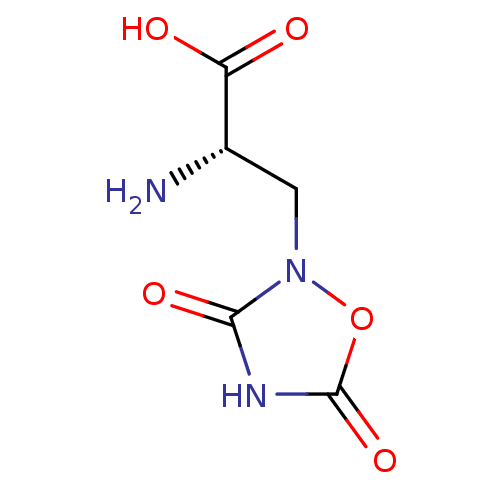
|
|---|


 Search and Browse
Search and Browse
 Download
Download
 Enter Data
Enter Data

 Schaffhauser, H; Richards, JG; Cartmell, J; Chaboz, S; Kemp, JA; Klingelschmidt, A; Messer, J; Stadler, H; Woltering, T; Mutel, V In vitro binding characteristics of a new selective group II metabotropic glutamate receptor radioligand, [3H]LY354740, in rat brain. Mol Pharmacol53:228-33 (1998) [PubMed]
Schaffhauser, H; Richards, JG; Cartmell, J; Chaboz, S; Kemp, JA; Klingelschmidt, A; Messer, J; Stadler, H; Woltering, T; Mutel, V In vitro binding characteristics of a new selective group II metabotropic glutamate receptor radioligand, [3H]LY354740, in rat brain. Mol Pharmacol53:228-33 (1998) [PubMed]