| Reaction Details |
|---|
 | Report a problem with these data |
| Target | Mitogen-activated protein kinase 9 |
|---|
| Ligand | BDBM16018 |
|---|
| Substrate/Competitor | n/a |
|---|
| Meas. Tech. | JNK Kinase Assay |
|---|
| IC50 | 110±0.0 nM |
|---|
| Citation |  Zhang, T; Inesta-Vaquera, F; Niepel, M; Zhang, J; Ficarro, SB; Machleidt, T; Xie, T; Marto, JA; Kim, N; Sim, T; Laughlin, JD; Park, H; LoGrasso, PV; Patricelli, M; Nomanbhoy, TK; Sorger, PK; Alessi, DR; Gray, NS Discovery of potent and selective covalent inhibitors of JNK. Chem Biol19:140-54 (2012) [PubMed] Article Zhang, T; Inesta-Vaquera, F; Niepel, M; Zhang, J; Ficarro, SB; Machleidt, T; Xie, T; Marto, JA; Kim, N; Sim, T; Laughlin, JD; Park, H; LoGrasso, PV; Patricelli, M; Nomanbhoy, TK; Sorger, PK; Alessi, DR; Gray, NS Discovery of potent and selective covalent inhibitors of JNK. Chem Biol19:140-54 (2012) [PubMed] Article |
|---|
| More Info.: | Get all data from this article, Assay Method |
|---|
| |
| Mitogen-activated protein kinase 9 |
|---|
| Name: | Mitogen-activated protein kinase 9 |
|---|
| Synonyms: | JNK-55 | JNK2 | JNK2/JNK3 | MAPK9 | MK09_HUMAN | Mitogen-Activated Protein Kinase 9 (JNK2) | Mitogen-activated protein kinase 8/9 | PRKM9 | SAPK1A | Stress-activated protein kinase JNK2 | c-Jun N-terminal kinase 2 | c-Jun N-terminal kinase 2 (JNK2) |
|---|
| Type: | Enzyme |
|---|
| Mol. Mass.: | 48131.49 |
|---|
| Organism: | Homo sapiens (Human) |
|---|
| Description: | JNK-2 was purchased from Upstate Cell Signaling Solutions (formerly Upstate Biotechnology). |
|---|
| Residue: | 424 |
|---|
| Sequence: | MSDSKCDSQFYSVQVADSTFTVLKRYQQLKPIGSGAQGIVCAAFDTVLGINVAVKKLSRP
FQNQTHAKRAYRELVLLKCVNHKNIISLLNVFTPQKTLEEFQDVYLVMELMDANLCQVIH
MELDHERMSYLLYQMLCGIKHLHSAGIIHRDLKPSNIVVKSDCTLKILDFGLARTACTNF
MMTPYVVTRYYRAPEVILGMGYKENVDIWSVGCIMGELVKGCVIFQGTDHIDQWNKVIEQ
LGTPSAEFMKKLQPTVRNYVENRPKYPGIKFEELFPDWIFPSESERDKIKTSQARDLLSK
MLVIDPDKRISVDEALRHPYITVWYDPAEAEAPPPQIYDAQLEEREHAIEEWKELIYKEV
MDWEERSKNGVVKDQPSDAAVSSNATPSQSSSINDISSMSTEQTLASDTDSSLDASTGPL
EGCR
|
|
|
|---|
| BDBM16018 |
|---|
| n/a |
|---|
| Name | BDBM16018 |
|---|
| Synonyms: | 14,15-diazatetracyclo[7.6.1.0^{2,7}.0^{13,16}]hexadeca-1(15),2(7),3,5,9(16),10,12-heptaen-8-one | CHEMBL7064 | ChemBiol10705 Compound 4 | JMC517015 Compound 2 | SP 600125 | SP-600125 | SP600125 | cid_8515 | dibenzo[cd,g]indazol-6(2H)-one |
|---|
| Type | Small organic molecule |
|---|
| Emp. Form. | C14H8N2O |
|---|
| Mol. Mass. | 220.2261 |
|---|
| SMILES | O=C1c2ccccc2-c2n[nH]c3cccc1c23 |
|---|
| Structure | 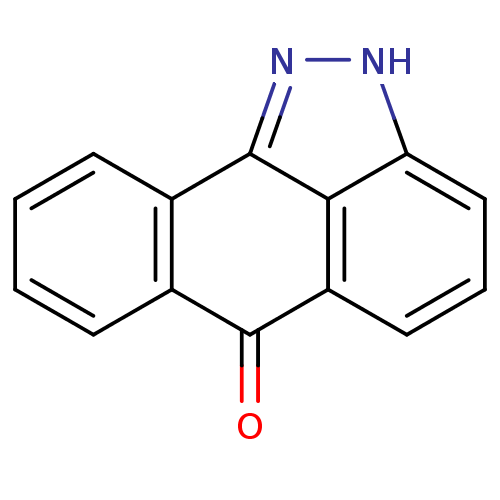
|
|---|


 Search and Browse
Search and Browse
 Download
Download
 Enter Data
Enter Data

 Zhang, T; Inesta-Vaquera, F; Niepel, M; Zhang, J; Ficarro, SB; Machleidt, T; Xie, T; Marto, JA; Kim, N; Sim, T; Laughlin, JD; Park, H; LoGrasso, PV; Patricelli, M; Nomanbhoy, TK; Sorger, PK; Alessi, DR; Gray, NS Discovery of potent and selective covalent inhibitors of JNK. Chem Biol19:140-54 (2012) [PubMed] Article
Zhang, T; Inesta-Vaquera, F; Niepel, M; Zhang, J; Ficarro, SB; Machleidt, T; Xie, T; Marto, JA; Kim, N; Sim, T; Laughlin, JD; Park, H; LoGrasso, PV; Patricelli, M; Nomanbhoy, TK; Sorger, PK; Alessi, DR; Gray, NS Discovery of potent and selective covalent inhibitors of JNK. Chem Biol19:140-54 (2012) [PubMed] Article