| Reaction Details |
|---|
 | Report a problem with these data |
| Target | Histone-lysine N-methyltransferase, H3 lysine-79 specific |
|---|
| Ligand | BDBM92649 |
|---|
| Substrate/Competitor | n/a |
|---|
| Meas. Tech. | Enzyme Assay |
|---|
| Ki | 0.3±0.0 nM |
|---|
| Kd | 0.10±0.02 nM |
|---|
| Citation |  Basavapathruni, A; Jin, L; Daigle, SR; Majer, CR; Therkelsen, CA; Wigle, TJ; Kuntz, KW; Chesworth, R; Pollock, RM; Scott, MP; Moyer, MP; Richon, VM; Copeland, RA; Olhava, EJ Conformational Adaptation Drives Potent, Selective and Durable Inhibition of the Human Protein Methyltransferase DOT1L. Chem Biol Drug Des80:971-80 (2012) [PubMed] Article Basavapathruni, A; Jin, L; Daigle, SR; Majer, CR; Therkelsen, CA; Wigle, TJ; Kuntz, KW; Chesworth, R; Pollock, RM; Scott, MP; Moyer, MP; Richon, VM; Copeland, RA; Olhava, EJ Conformational Adaptation Drives Potent, Selective and Durable Inhibition of the Human Protein Methyltransferase DOT1L. Chem Biol Drug Des80:971-80 (2012) [PubMed] Article |
|---|
| More Info.: | Get all data from this article, Assay Method |
|---|
| |
| Histone-lysine N-methyltransferase, H3 lysine-79 specific |
|---|
| Name: | Histone-lysine N-methyltransferase, H3 lysine-79 specific |
|---|
| Synonyms: | 2.1.1.43 | DOT1-like protein | DOT1-like protein (Dot1L) | DOT1L | DOT1L_HUMAN | H3-K79-HMTase | Histone H3-K79 methyltransferase | Histone H3-K79 methyltransferase (DOT1L) | Histone Methyltransferase DOT1L | Histone-lysine N-methyltransferase, H3 lysine-79 specific (DOT1L) | KIAA1814 | KMT4 | Lysine N-methyltransferase 4 |
|---|
| Type: | Protein |
|---|
| Mol. Mass.: | 184911.91 |
|---|
| Organism: | Homo sapiens (Human) |
|---|
| Description: | Q8TEK3 |
|---|
| Residue: | 1537 |
|---|
| Sequence: | MGEKLELRLKSPVGAEPAVYPWPLPVYDKHHDAAHEIIETIRWVCEEIPDLKLAMENYVL
IDYDTKSFESMQRLCDKYNRAIDSIHQLWKGTTQPMKLNTRPSTGLLRHILQQVYNHSVT
DPEKLNNYEPFSPEVYGETSFDLVAQMIDEIKMTDDDLFVDLGSGVGQVVLQVAAATNCK
HHYGVEKADIPAKYAETMDREFRKWMKWYGKKHAEYTLERGDFLSEEWRERIANTSVIFV
NNFAFGPEVDHQLKERFANMKEGGRIVSSKPFAPLNFRINSRNLSDIGTIMRVVELSPLK
GSVSWTGKPVSYYLHTIDRTILENYFSSLKNPKLREEQEAARRRQQRESKSNAATPTKGP
EGKVAGPADAPMDSGAEEEKAGAATVKKPSPSKARKKKLNKKGRKMAGRKRGRPKKMNTA
NPERKPKKNQTALDALHAQTVSQTAASSPQDAYRSPHSPFYQLPPSVQRHSPNPLLVAPT
PPALQKLLESFKIQYLQFLAYTKTPQYKASLQELLGQEKEKNAQLLGAAQQLLSHCQAQK
EEIRRLFQQKLDELGVKALTYNDLIQAQKEISAHNQQLREQSEQLEQDNRALRGQSLQLL
KARCEELQLDWATLSLEKLLKEKQALKSQISEKQRHCLELQISIVELEKSQRQQELLQLK
SCVPPDDALSLHLRGKGALGRELEPDASRLHLELDCTKFSLPHLSSMSPELSMNGQAAGY
ELCGVLSRPSSKQNTPQYLASPLDQEVVPCTPSHVGRPRLEKLSGLAAPDYTRLSPAKIV
LRRHLSQDHTVPGRPAASELHSRAEHTKENGLPYQSPSVPGSMKLSPQDPRPLSPGALQL
AGEKSSEKGLRERAYGSSGELITSLPISIPLSTVQPNKLPVSIPLASVVLPSRAERARST
PSPVLQPRDPSSTLEKQIGANAHGAGSRSLALAPAGFSYAGSVAISGALAGSPASLTPGA
EPATLDESSSSGSLFATVGSRSSTPQHPLLLAQPRNSLPASPAHQLSSSPRLGGAAQGPL
PEASKGDLPSDSGFSDPESEAKRRIVFTITTGAGSAKQSPSSKHSPLTASARGDCVPSHG
QDSRRRGRRKRASAGTPSLSAGVSPKRRALPSVAGLFTQPSGSPLNLNSMVSNINQPLEI
TAISSPETSLKSSPVPYQDHDQPPVLKKERPLSQTNGAHYSPLTSDEEPGSEDEPSSARI
ERKIATISLESKSPPKTLENGGGLAGRKPAPAGEPVNSSKWKSTFSPISDIGLAKSADSP
LQASSALSQNSLFTFRPALEEPSADAKLAAHPRKGFPGSLSGADGLSPGTNPANGCTFGG
GLAADLSLHSFSDGASLPHKGPEAAGLSSPLSFPSQRGKEGSDANPFLSKRQLDGLAGLK
GEGSRGKEAGEGGLPLCGPTDKTPLLSGKAAKARDREVDLKNGHNLFISAAAVPPGSLLS
GPGLAPAASSAGGAASSAQTHRSFLGPFPPGPQFALGPMSLQANLGSVAGSSVLQSLFSS
VPAAAGLVHVSSAATRLTNSHAMGSFSGVAGGTVGGN
|
|
|
|---|
| BDBM92649 |
|---|
| n/a |
|---|
| Name | BDBM92649 |
|---|
| Synonyms: | EPZ004777 |
|---|
| Type | Organic molecule |
|---|
| Emp. Form. | C28H41N7O4 |
|---|
| Mol. Mass. | 539.6696 |
|---|
| SMILES | CC(C)N(CCCNC(=O)Nc1ccc(cc1)C(C)(C)C)CC1OC([C@H](O)[C@@H]1O)n1ccc2c(N)ncnc12 |r| |
|---|
| Structure | 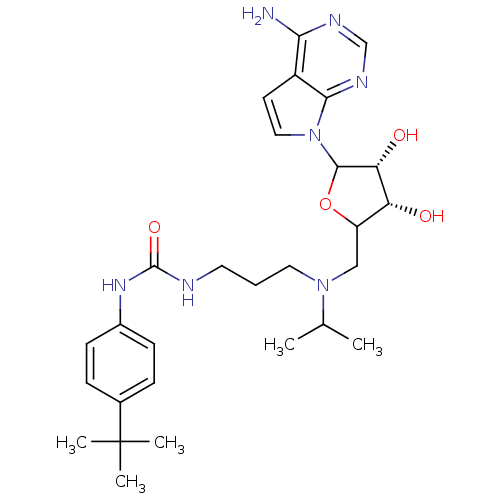
|
|---|


 Search and Browse
Search and Browse
 Download
Download
 Enter Data
Enter Data

 Basavapathruni, A; Jin, L; Daigle, SR; Majer, CR; Therkelsen, CA; Wigle, TJ; Kuntz, KW; Chesworth, R; Pollock, RM; Scott, MP; Moyer, MP; Richon, VM; Copeland, RA; Olhava, EJ Conformational Adaptation Drives Potent, Selective and Durable Inhibition of the Human Protein Methyltransferase DOT1L. Chem Biol Drug Des80:971-80 (2012) [PubMed] Article
Basavapathruni, A; Jin, L; Daigle, SR; Majer, CR; Therkelsen, CA; Wigle, TJ; Kuntz, KW; Chesworth, R; Pollock, RM; Scott, MP; Moyer, MP; Richon, VM; Copeland, RA; Olhava, EJ Conformational Adaptation Drives Potent, Selective and Durable Inhibition of the Human Protein Methyltransferase DOT1L. Chem Biol Drug Des80:971-80 (2012) [PubMed] Article