| Reaction Details |
|---|
 | Report a problem with these data |
| Target | Beta-galactosidase |
|---|
| Ligand | BDBM163641 |
|---|
| Substrate/Competitor | n/a |
|---|
| Meas. Tech. | Inhibition Assays |
|---|
| Ki | 2.8e+3± 3e+2 nM |
|---|
| Citation |  Wallace, BD; Roberts, AB; Pollet, RM; Ingle, JD; Biernat, KA; Pellock, SJ; Venkatesh, MK; Guthrie, L; O'Neal, SK; Robinson, SJ; Dollinger, M; Figueroa, E; McShane, SR; Cohen, RD; Jin, J; Frye, SV; Zamboni, WC; Pepe-Ranney, C; Mani, S; Kelly, L; Redinbo, MR Structure and Inhibition of Microbiome �-Glucuronidases Essential to the Alleviation of Cancer Drug Toxicity. Chem Biol22:1238-49 (2015) [PubMed] Article Wallace, BD; Roberts, AB; Pollet, RM; Ingle, JD; Biernat, KA; Pellock, SJ; Venkatesh, MK; Guthrie, L; O'Neal, SK; Robinson, SJ; Dollinger, M; Figueroa, E; McShane, SR; Cohen, RD; Jin, J; Frye, SV; Zamboni, WC; Pepe-Ranney, C; Mani, S; Kelly, L; Redinbo, MR Structure and Inhibition of Microbiome �-Glucuronidases Essential to the Alleviation of Cancer Drug Toxicity. Chem Biol22:1238-49 (2015) [PubMed] Article |
|---|
| More Info.: | Get all data from this article, Assay Method |
|---|
| |
| Beta-galactosidase |
|---|
| Name: | Beta-galactosidase |
|---|
| Synonyms: | Beta-glucuronidase (SaGUS) |
|---|
| Type: | Protein |
|---|
| Mol. Mass.: | 69235.31 |
|---|
| Organism: | Streptococcus agalactiae (Firmicutes) |
|---|
| Description: | A0A0E1EGE5 |
|---|
| Residue: | 599 |
|---|
| Sequence: | MLYPLLTKTRNTYDLGGIWNFKLGEHNPNELLPSDEVMVIPTSFNDLMVSKEKRDYIGDF
WYEKVIEVPKVSEGEEMVLRFGSVTHQAKIYVDGILVGEHKGGFTPFEVLVPECKYNNEK
IKVSICANNVLDYTTLPVGNYSEIIQEDGSIKKKVRENFDFFNYAGVHRPLKLMIRPKNH
ISDITITSRLSDDLQSADLHFLVETNQKVDEVRISVFDEDNKLVGETKDSRLFLSDVHLW
EVLNAYLYTARVEIFVDNQLQDVYEENFGLREIEVTNGQFLLNRKPIYFKGFGKHEDTFI
NGRGLNEAANLMDLNLLKDIGANSFRTSHYPYSEEMMRLADRMGVLVIDEVPAVGLFQNF
NASLDLSPKDNGTWSLMQTKAAHEQAIQELVKRDKNHPSVVMWVVANEPASHEAGAHDYF
EPLVKLYKDLDPQKRPVTLVNILMATPDRDQVMDLVDVVCLNRYYGWYVDHGDLTNAEVG
LRKELLEWQDKFPDKPIIITEYGADTLPGLHSTWNIPYTEEFQCDFYEMSHRVFDGIPNL
VGEQVWNFADFETNLMILRVQGNHKGLFSRNRQPKQVVKEFKKRWMTIPHYHNKKNSVK
|
|
|
|---|
| BDBM163641 |
|---|
| n/a |
|---|
| Name | BDBM163641 |
|---|
| Synonyms: | 1[2(3Fluorophenyl)5H,6H,7H,8Hpyrazolo[3,2b]quinazolin9yl]piperazine (Inh 6) | US9334288, 7 |
|---|
| Type | Small organic molecule |
|---|
| Emp. Form. | C20H22FN5 |
|---|
| Mol. Mass. | 351.4206 |
|---|
| SMILES | Fc1cccc(c1)-c1cc2nc3CCCCc3c(N3CCNCC3)n2n1 |
|---|
| Structure | 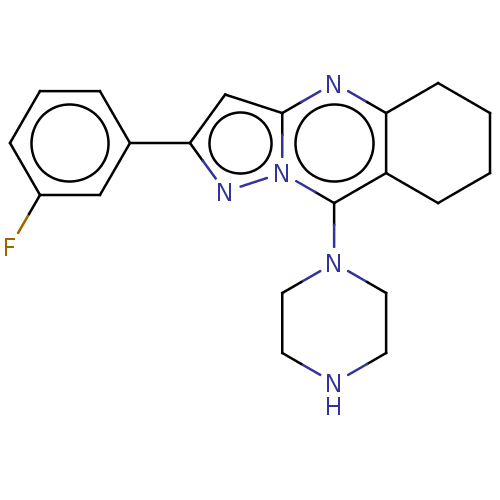
|
|---|


 Search and Browse
Search and Browse
 Download
Download
 Enter Data
Enter Data

 Wallace, BD; Roberts, AB; Pollet, RM; Ingle, JD; Biernat, KA; Pellock, SJ; Venkatesh, MK; Guthrie, L; O'Neal, SK; Robinson, SJ; Dollinger, M; Figueroa, E; McShane, SR; Cohen, RD; Jin, J; Frye, SV; Zamboni, WC; Pepe-Ranney, C; Mani, S; Kelly, L; Redinbo, MR Structure and Inhibition of Microbiome �-Glucuronidases Essential to the Alleviation of Cancer Drug Toxicity. Chem Biol22:1238-49 (2015) [PubMed] Article
Wallace, BD; Roberts, AB; Pollet, RM; Ingle, JD; Biernat, KA; Pellock, SJ; Venkatesh, MK; Guthrie, L; O'Neal, SK; Robinson, SJ; Dollinger, M; Figueroa, E; McShane, SR; Cohen, RD; Jin, J; Frye, SV; Zamboni, WC; Pepe-Ranney, C; Mani, S; Kelly, L; Redinbo, MR Structure and Inhibition of Microbiome �-Glucuronidases Essential to the Alleviation of Cancer Drug Toxicity. Chem Biol22:1238-49 (2015) [PubMed] Article