| Reaction Details |
|---|
 | Report a problem with these data |
| Target | Histone deacetylase 3 |
|---|
| Ligand | BDBM19149 |
|---|
| Substrate/Competitor | n/a |
|---|
| Meas. Tech. | In-vitro HDAC Enzymatic Endpoint Assay |
|---|
| pH | 7.4±n/a |
|---|
| Temperature | 298.15±n/a K |
|---|
| IC50 | 3± 0.5 nM |
|---|
| Comments | extracted |
|---|
| Citation |  Wagner, FF; Lundh, M; Kaya, T; McCarren, P; Zhang, YL; Chattopadhyay, S; Gale, JP; Galbo, T; Fisher, SL; Meier, BC; Vetere, A; Richardson, S; Morgan, NG; Christensen, DP; Gilbert, TJ; Hooker, JM; Leroy, M; Walpita, D; Mandrup-Poulsen, T; Wagner, BK; Holson, EB An Isochemogenic Set of Inhibitors To Define the Therapeutic Potential of Histone Deacetylases in �-Cell Protection. ACS Chem Biol11:363-74 (2016) [PubMed] Article Wagner, FF; Lundh, M; Kaya, T; McCarren, P; Zhang, YL; Chattopadhyay, S; Gale, JP; Galbo, T; Fisher, SL; Meier, BC; Vetere, A; Richardson, S; Morgan, NG; Christensen, DP; Gilbert, TJ; Hooker, JM; Leroy, M; Walpita, D; Mandrup-Poulsen, T; Wagner, BK; Holson, EB An Isochemogenic Set of Inhibitors To Define the Therapeutic Potential of Histone Deacetylases in �-Cell Protection. ACS Chem Biol11:363-74 (2016) [PubMed] Article |
|---|
| More Info.: | Get all data from this article, Assay Method |
|---|
| |
| Histone deacetylase 3 |
|---|
| Name: | Histone deacetylase 3 |
|---|
| Synonyms: | HD3 | HDAC3 | HDAC3_HUMAN | Histone deacetylase 3 (HDAC3) | Human HDAC3 | RPD3-2 | SMAP45 |
|---|
| Type: | Enzyme |
|---|
| Mol. Mass.: | 48829.55 |
|---|
| Organism: | Homo sapiens (Human) |
|---|
| Description: | O15379 |
|---|
| Residue: | 428 |
|---|
| Sequence: | MAKTVAYFYDPDVGNFHYGAGHPMKPHRLALTHSLVLHYGLYKKMIVFKPYQASQHDMCR
FHSEDYIDFLQRVSPTNMQGFTKSLNAFNVGDDCPVFPGLFEFCSRYTGASLQGATQLNN
KICDIAINWAGGLHHAKKFEASGFCYVNDIVIGILELLKYHPRVLYIDIDIHHGDGVQEA
FYLTDRVMTVSFHKYGNYFFPGTGDMYEVGAESGRYYCLNVPLRDGIDDQSYKHLFQPVI
NQVVDFYQPTCIVLQCGADSLGCDRLGCFNLSIRGHGECVEYVKSFNIPLLVLGGGGYTV
RNVARCWTYETSLLVEEAISEELPYSEYFEYFAPDFTLHPDVSTRIENQNSRQYLDQIRQ
TIFENLKMLNHAPSVQIHDVPADLLTYDRTDEADAEERGPEENYSRPEAPNEFYDGDHDN
DKESDVEI
|
|
|
|---|
| BDBM19149 |
|---|
| n/a |
|---|
| Name | BDBM19149 |
|---|
| Synonyms: | CHEMBL98 | N-hydroxy-N'-phenyloctanediamide | SAHA | US10011611, SAHA | US10188756, Compound SAHA | US11207431, SAHA | US11505523, Compound SAHA | US9115116, SAHA | US9353061, SAHA | US9428447, SAHA | US9695181, Vorinostat | Vorinostat | Zolinza | cid_5311 | suberoylanilide hydroxamic acid |
|---|
| Type | Small organic molecule |
|---|
| Emp. Form. | C14H20N2O3 |
|---|
| Mol. Mass. | 264.3202 |
|---|
| SMILES | ONC(=O)CCCCCCC(=O)Nc1ccccc1 |
|---|
| Structure | 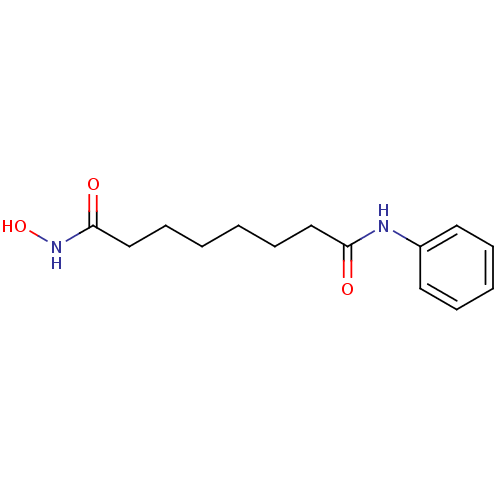
|
|---|


 Search and Browse
Search and Browse
 Download
Download
 Enter Data
Enter Data

 Wagner, FF; Lundh, M; Kaya, T; McCarren, P; Zhang, YL; Chattopadhyay, S; Gale, JP; Galbo, T; Fisher, SL; Meier, BC; Vetere, A; Richardson, S; Morgan, NG; Christensen, DP; Gilbert, TJ; Hooker, JM; Leroy, M; Walpita, D; Mandrup-Poulsen, T; Wagner, BK; Holson, EB An Isochemogenic Set of Inhibitors To Define the Therapeutic Potential of Histone Deacetylases in �-Cell Protection. ACS Chem Biol11:363-74 (2016) [PubMed] Article
Wagner, FF; Lundh, M; Kaya, T; McCarren, P; Zhang, YL; Chattopadhyay, S; Gale, JP; Galbo, T; Fisher, SL; Meier, BC; Vetere, A; Richardson, S; Morgan, NG; Christensen, DP; Gilbert, TJ; Hooker, JM; Leroy, M; Walpita, D; Mandrup-Poulsen, T; Wagner, BK; Holson, EB An Isochemogenic Set of Inhibitors To Define the Therapeutic Potential of Histone Deacetylases in �-Cell Protection. ACS Chem Biol11:363-74 (2016) [PubMed] Article