| Reaction Details |
|---|
 | Report a problem with these data |
| Target | Lysine-specific demethylase 5A [1-588] |
|---|
| Ligand | BDBM191602 |
|---|
| Substrate/Competitor | n/a |
|---|
| Meas. Tech. | Isothermal Titration Calorimetry (ITC) |
|---|
| Kd | 4.5e+2± 3e+1 nM |
|---|
| Citation |  Horton, JR; Liu, X; Gale, M; Wu, L; Shanks, JR; Zhang, X; Webber, PJ; Bell, JS; Kales, SC; Mott, BT; Rai, G; Jansen, DJ; Henderson, MJ; Urban, DJ; Hall, MD; Simeonov, A; Maloney, DJ; Johns, MA; Fu, H; Jadhav, A; Vertino, PM; Yan, Q; Cheng, X Structural Basis for KDM5A Histone Lysine Demethylase Inhibition by Diverse Compounds. Cell Chem Biol23:769-81 (2016) [PubMed] Article Horton, JR; Liu, X; Gale, M; Wu, L; Shanks, JR; Zhang, X; Webber, PJ; Bell, JS; Kales, SC; Mott, BT; Rai, G; Jansen, DJ; Henderson, MJ; Urban, DJ; Hall, MD; Simeonov, A; Maloney, DJ; Johns, MA; Fu, H; Jadhav, A; Vertino, PM; Yan, Q; Cheng, X Structural Basis for KDM5A Histone Lysine Demethylase Inhibition by Diverse Compounds. Cell Chem Biol23:769-81 (2016) [PubMed] Article |
|---|
| More Info.: | Get all data from this article, Assay Method |
|---|
| |
| Lysine-specific demethylase 5A [1-588] |
|---|
| Name: | Lysine-specific demethylase 5A [1-588] |
|---|
| Synonyms: | JARID1A | KDM5A | KDM5A_HUMAN | Lysine-specific demethylase 5A (KDM5A(aa 1-588)-AP) | RBBP2 | RBP2 |
|---|
| Type: | Protein |
|---|
| Mol. Mass.: | 66955.44 |
|---|
| Organism: | Homo sapiens (Human) |
|---|
| Description: | Truncation 1-588 aa. Contains deletion of ARID and PhD1 domains. |
|---|
| Residue: | 588 |
|---|
| Sequence: | MAGVGPGGYAAEFVPPPECPVFEPSWEEFTDPLSFIGRIRPLAEKTGICKIRPPKDWQPP
FACEVKSFRFTPRVQRLNELEAMTRVRLDFLDQLAKFWELQGSTLKIPVVERKILDLYAL
SKIVASKGGFEMVTKEKKWSKVGSRLGYLPGKGTGSLLKSHYERILYPYELFQSGVSLMG
VQMPNLDLKEKVEPEVLSTDTQTSPEPGTRMNILPKRTRRVKTQSESGDVSRNTELKKLQ
IFGAGPKVVGLAMGTKDKEDEVTRRRKVTNRSDAFNMQMRQRKGTLSVNFVDLYVCMFCG
RGNNEDKLLLCDGCDDSYHTFCLIPPLPDVPKGDWRCPKCVAEECSKPREAFGFEQAVRE
YTLQSFGEMADNFKSDYFNMPVHMVPTELVEKEFWRLVSSIEEDVIVEYGADISSKDFGS
GFPVKDGRRKILPEEEEYALSGWNLNNMPVLEQSVLAHINVDISGMKVPWLYVGMCFSSF
CWHIEDHWSYSINYLHWGEPKTWYGVPSHAAEQLEEVMRELAPELFESQPDLLHQLVTIM
NPNVLMEHGVPVYRTNQCAGEFVVTFPRAYHSGFNQGYNFAEAVNFCT
|
|
|
|---|
| BDBM191602 |
|---|
| n/a |
|---|
| Name | BDBM191602 |
|---|
| Synonyms: | 3-((1-methyl-1Hpyrrolo[2,3-b]pyridin-3-yl)amino)isonicotinic acid (N12) | US10040779, Example 52 | US10336727, Example 52 | US9617242, Example 52 |
|---|
| Type | Small organic molecule |
|---|
| Emp. Form. | C14H12N4O2 |
|---|
| Mol. Mass. | 268.2707 |
|---|
| SMILES | Cn1cc(Nc2cnccc2C(O)=O)c2cccnc12 |
|---|
| Structure | 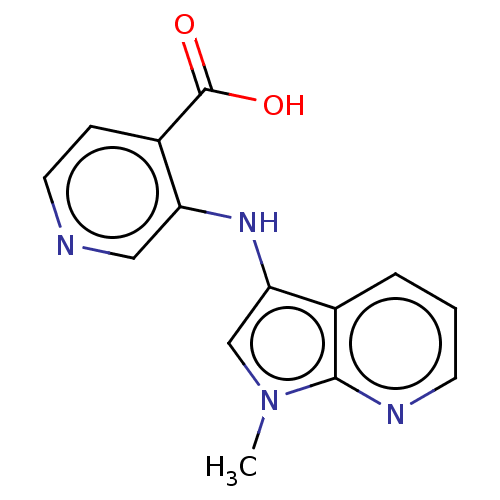
|
|---|


 Search and Browse
Search and Browse
 Download
Download
 Enter Data
Enter Data

 Horton, JR; Liu, X; Gale, M; Wu, L; Shanks, JR; Zhang, X; Webber, PJ; Bell, JS; Kales, SC; Mott, BT; Rai, G; Jansen, DJ; Henderson, MJ; Urban, DJ; Hall, MD; Simeonov, A; Maloney, DJ; Johns, MA; Fu, H; Jadhav, A; Vertino, PM; Yan, Q; Cheng, X Structural Basis for KDM5A Histone Lysine Demethylase Inhibition by Diverse Compounds. Cell Chem Biol23:769-81 (2016) [PubMed] Article
Horton, JR; Liu, X; Gale, M; Wu, L; Shanks, JR; Zhang, X; Webber, PJ; Bell, JS; Kales, SC; Mott, BT; Rai, G; Jansen, DJ; Henderson, MJ; Urban, DJ; Hall, MD; Simeonov, A; Maloney, DJ; Johns, MA; Fu, H; Jadhav, A; Vertino, PM; Yan, Q; Cheng, X Structural Basis for KDM5A Histone Lysine Demethylase Inhibition by Diverse Compounds. Cell Chem Biol23:769-81 (2016) [PubMed] Article