| Reaction Details |
|---|
 | Report a problem with these data |
| Target | Lysine-specific demethylase 6B |
|---|
| Ligand | BDBM26106 |
|---|
| Substrate/Competitor | n/a |
|---|
| Meas. Tech. | Demethylase AlphaScreen Assay |
|---|
| pH | 7.5±n/a |
|---|
| IC50 | 6.8e+2±n/a nM |
|---|
| Comments | extracted |
|---|
| Citation |  Johansson, C; Velupillai, S; Tumber, A; Szykowska, A; Hookway, ES; Nowak, RP; Strain-Damerell, C; Gileadi, C; Philpott, M; Burgess-Brown, N; Wu, N; Kopec, J; Nuzzi, A; Steuber, H; Egner, U; Badock, V; Munro, S; LaThangue, NB; Westaway, S; Brown, J; Athanasou, N; Prinjha, R; Brennan, PE; Oppermann, U Structural analysis of human KDM5B guides histone demethylase inhibitor development. Nat Chem Biol12:539-45 (2016) [PubMed] Article Johansson, C; Velupillai, S; Tumber, A; Szykowska, A; Hookway, ES; Nowak, RP; Strain-Damerell, C; Gileadi, C; Philpott, M; Burgess-Brown, N; Wu, N; Kopec, J; Nuzzi, A; Steuber, H; Egner, U; Badock, V; Munro, S; LaThangue, NB; Westaway, S; Brown, J; Athanasou, N; Prinjha, R; Brennan, PE; Oppermann, U Structural analysis of human KDM5B guides histone demethylase inhibitor development. Nat Chem Biol12:539-45 (2016) [PubMed] Article |
|---|
| More Info.: | Get all data from this article, Assay Method |
|---|
| |
| Lysine-specific demethylase 6B |
|---|
| Name: | Lysine-specific demethylase 6B |
|---|
| Synonyms: | JMJD3 | JmjC domain-containing protein 3 | Jumonji domain-containing protein 3 | KDM6B | KDM6B_HUMAN | KIAA0346 | Lysine demethylase 6B | Lysine-specific demethylase 6B | Lysine-specific demethylase 6B (KDM6B(JMJD3)) | Lysine-specific demethylase 6B (KDM6B) |
|---|
| Type: | Protein |
|---|
| Mol. Mass.: | 176672.55 |
|---|
| Organism: | Homo sapiens (Human) |
|---|
| Description: | O15054 |
|---|
| Residue: | 1643 |
|---|
| Sequence: | MHRAVDPPGARAAREAFALGGLSCAGAWSSCPPHPPPRSAWLPGGRCSASIGQPPLPAPL
PPSHGSSSGHPSKPYYAPGAPTPRPLHGKLESLHGCVQALLREPAQPGLWEQLGQLYESE
HDSEEATRCYHSALRYGGSFAELGPRIGRLQQAQLWNFHTGSCQHRAKVLPPLEQVWNLL
HLEHKRNYGAKRGGPPVKRAAEPPVVQPVPPAALSGPSGEEGLSPGGKRRRGCNSEQTGL
PPGLPLPPPPLPPPPPPPPPPPPPLPGLATSPPFQLTKPGLWSTLHGDAWGPERKGSAPP
ERQEQRHSLPHPYPYPAPAYTAHPPGHRLVPAAPPGPGPRPPGAESHGCLPATRPPGSDL
RESRVQRSRMDSSVSPAATTACVPYAPSRPPGLPGTTTSSSSSSSSNTGLRGVEPNPGIP
GADHYQTPALEVSHHGRLGPSAHSSRKPFLGAPAATPHLSLPPGPSSPPPPPCPRLLRPP
PPPAWLKGPACRAAREDGEILEELFFGTEGPPRPAPPPLPHREGFLGPPASRFSVGTQDS
HTPPTPPTPTTSSSNSNSGSHSSSPAGPVSFPPPPYLARSIDPLPRPPSPAQNPQDPPLV
PLTLALPPAPPSSCHQNTSGSFRRPESPRPRVSFPKTPEVGPGPPPGPLSKAPQPVPPGV
GELPARGPRLFDFPPTPLEDQFEEPAEFKILPDGLANIMKMLDESIRKEEEQQQHEAGVA
PQPPLKEPFASLQSPFPTDTAPTTTAPAVAVTTTTTTTTTTTATQEEEKKPPPALPPPPP
LAKFPPPSQPQPPPPPPPSPASLLKSLASVLEGQKYCYRGTGAAVSTRPGPLPTTQYSPG
PPSGATALPPTSAAPSAQGSPQPSASSSSQFSTSGGPWARERRAGEEPVPGPMTPTQPPP
PLSLPPARSESEVLEEISRACETLVERVGRSATDPADPVDTAEPADSGTERLLPPAQAKE
EAGGVAAVSGSCKRRQKEHQKEHRRHRRACKDSVGRRPREGRAKAKAKVPKEKSRRVLGN
LDLQSEEIQGREKSRPDLGGASKAKPPTAPAPPSAPAPSAQPTPPSASVPGKKAREEAPG
PPGVSRADMLKLRSLSEGPPKELKIRLIKVESGDKETFIASEVEERRLRMADLTISHCAA
DVVRASRNAKVKGKFRESYLSPAQSVKPKINTEEKLPREKLNPPTPSIYLESKRDAFSPV
LLQFCTDPRNPITVIRGLAGSLRLNLGLFSTKTLVEASGEHTVEVRTQVQQPSDENWDLT
GTRQIWPCESSRSHTTIAKYAQYQASSFQESLQEEKESEDEESEEPDSTTGTPPSSAPDP
KNHHIIKFGTNIDLSDAKRWKPQLQELLKLPAFMRVTSTGNMLSHVGHTILGMNTVQLYM
KVPGSRTPGHQENNNFCSVNINIGPGDCEWFAVHEHYWETISAFCDRHGVDYLTGSWWPI
LDDLYASNIPVYRFVQRPGDLVWINAGTVHWVQATGWCNNIAWNVGPLTAYQYQLALERY
EWNEVKNVKSIVPMIHVSWNVARTVKISDPDLFKMIKFCLLQSMKHCQVQRESLVRAGKK
IAYQGRVKDEPAYYCNECDVEVFNILFVTSENGSRNTYLVHCEGCARRRSAGLQGVVVLE
QYRTEELAQAYDAFTLAPASTSR
|
|
|
|---|
| BDBM26106 |
|---|
| n/a |
|---|
| Name | BDBM26106 |
|---|
| Synonyms: | CHEMBL90852 | N-oxalyl glycine, 1a | NOG | Oxalylglycine | [(carboxymethyl)carbamoyl]formic acid |
|---|
| Type | Small organic molecule |
|---|
| Emp. Form. | C4H5NO5 |
|---|
| Mol. Mass. | 147.0862 |
|---|
| SMILES | OC(=O)CNC(=O)C(O)=O |
|---|
| Structure | 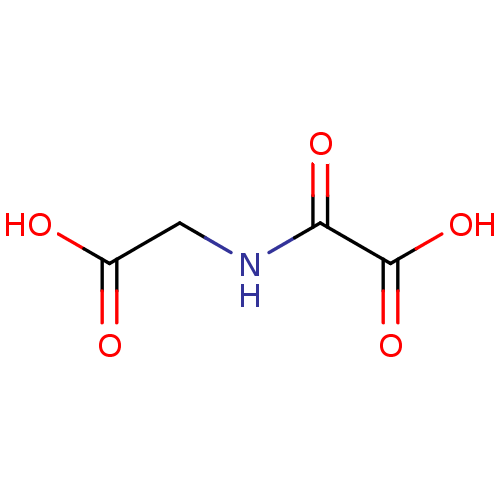
|
|---|


 Search and Browse
Search and Browse
 Download
Download
 Enter Data
Enter Data

 Johansson, C; Velupillai, S; Tumber, A; Szykowska, A; Hookway, ES; Nowak, RP; Strain-Damerell, C; Gileadi, C; Philpott, M; Burgess-Brown, N; Wu, N; Kopec, J; Nuzzi, A; Steuber, H; Egner, U; Badock, V; Munro, S; LaThangue, NB; Westaway, S; Brown, J; Athanasou, N; Prinjha, R; Brennan, PE; Oppermann, U Structural analysis of human KDM5B guides histone demethylase inhibitor development. Nat Chem Biol12:539-45 (2016) [PubMed] Article
Johansson, C; Velupillai, S; Tumber, A; Szykowska, A; Hookway, ES; Nowak, RP; Strain-Damerell, C; Gileadi, C; Philpott, M; Burgess-Brown, N; Wu, N; Kopec, J; Nuzzi, A; Steuber, H; Egner, U; Badock, V; Munro, S; LaThangue, NB; Westaway, S; Brown, J; Athanasou, N; Prinjha, R; Brennan, PE; Oppermann, U Structural analysis of human KDM5B guides histone demethylase inhibitor development. Nat Chem Biol12:539-45 (2016) [PubMed] Article