| Reaction Details |
|---|
 | Report a problem with these data |
| Target | Histone deacetylase 1 |
|---|
| Ligand | BDBM19410 |
|---|
| Substrate/Competitor | n/a |
|---|
| Meas. Tech. | Enzymatic HDAC Activity Assay |
|---|
| pH | 7.5±n/a |
|---|
| IC50 | 243±n/a nM |
|---|
| Comments | extracted |
|---|
| Citation |  Lauffer, BE; Mintzer, R; Fong, R; Mukund, S; Tam, C; Zilberleyb, I; Flicke, B; Ritscher, A; Fedorowicz, G; Vallero, R; Ortwine, DF; Gunzner, J; Modrusan, Z; Neumann, L; Koth, CM; Lupardus, PJ; Kaminker, JS; Heise, CE; Steiner, P Histone deacetylase (HDAC) inhibitor kinetic rate constants correlate with cellular histone acetylation but not transcription and cell viability. J Biol Chem288:26926-43 (2013) [PubMed] Article Lauffer, BE; Mintzer, R; Fong, R; Mukund, S; Tam, C; Zilberleyb, I; Flicke, B; Ritscher, A; Fedorowicz, G; Vallero, R; Ortwine, DF; Gunzner, J; Modrusan, Z; Neumann, L; Koth, CM; Lupardus, PJ; Kaminker, JS; Heise, CE; Steiner, P Histone deacetylase (HDAC) inhibitor kinetic rate constants correlate with cellular histone acetylation but not transcription and cell viability. J Biol Chem288:26926-43 (2013) [PubMed] Article |
|---|
| More Info.: | Get all data from this article, Assay Method |
|---|
| |
| Histone deacetylase 1 |
|---|
| Name: | Histone deacetylase 1 |
|---|
| Synonyms: | Cereblon/Histone deacetylase 1 | HD1 | HDAC1 | HDAC1_HUMAN | Histone deacetylase 1 (HDAC1) | Human HDAC1 | RPD3L1 |
|---|
| Type: | Enzyme |
|---|
| Mol. Mass.: | 55090.27 |
|---|
| Organism: | Homo sapiens (Human) |
|---|
| Description: | Q13547 |
|---|
| Residue: | 482 |
|---|
| Sequence: | MAQTQGTRRKVCYYYDGDVGNYYYGQGHPMKPHRIRMTHNLLLNYGLYRKMEIYRPHKAN
AEEMTKYHSDDYIKFLRSIRPDNMSEYSKQMQRFNVGEDCPVFDGLFEFCQLSTGGSVAS
AVKLNKQQTDIAVNWAGGLHHAKKSEASGFCYVNDIVLAILELLKYHQRVLYIDIDIHHG
DGVEEAFYTTDRVMTVSFHKYGEYFPGTGDLRDIGAGKGKYYAVNYPLRDGIDDESYEAI
FKPVMSKVMEMFQPSAVVLQCGSDSLSGDRLGCFNLTIKGHAKCVEFVKSFNLPMLMLGG
GGYTIRNVARCWTYETAVALDTEIPNELPYNDYFEYFGPDFKLHISPSNMTNQNTNEYLE
KIKQRLFENLRMLPHAPGVQMQAIPEDAIPEESGDEDEDDPDKRISICSSDKRIACEEEF
SDSEEEGEGGRKNSSNFKKAKRVKTEDEKEKDPEEKKEVTEEEKTKEEKPEAKGVKEEVK
LA
|
|
|
|---|
| BDBM19410 |
|---|
| n/a |
|---|
| Name | BDBM19410 |
|---|
| Synonyms: | CHEMBL27759 | MS-275 | US11377423, MS-275 | US11672788, Compound Entinostat | US9265734, MS-275 | US9796664, Compound MS-275 | benzamide-type inhibitor, 3 | pyridin-3-ylmethyl N-({4-[(2-aminophenyl)carbamoyl]phenyl}methyl)carbamate |
|---|
| Type | Small organic molecule |
|---|
| Emp. Form. | C21H20N4O3 |
|---|
| Mol. Mass. | 376.4085 |
|---|
| SMILES | Nc1ccccc1NC(=O)c1ccc(CNC(=O)OCc2cccnc2)cc1 |
|---|
| Structure | 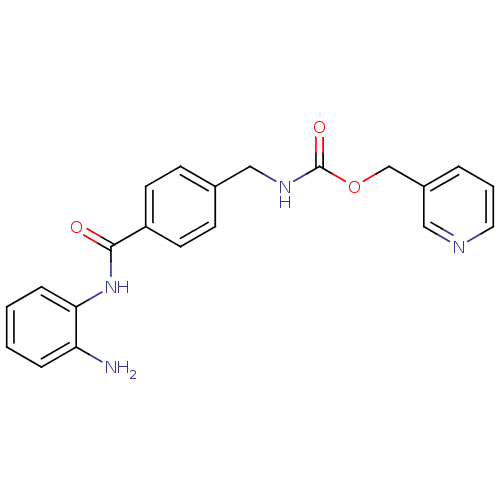
|
|---|


 Search and Browse
Search and Browse
 Download
Download
 Enter Data
Enter Data

 Lauffer, BE; Mintzer, R; Fong, R; Mukund, S; Tam, C; Zilberleyb, I; Flicke, B; Ritscher, A; Fedorowicz, G; Vallero, R; Ortwine, DF; Gunzner, J; Modrusan, Z; Neumann, L; Koth, CM; Lupardus, PJ; Kaminker, JS; Heise, CE; Steiner, P Histone deacetylase (HDAC) inhibitor kinetic rate constants correlate with cellular histone acetylation but not transcription and cell viability. J Biol Chem288:26926-43 (2013) [PubMed] Article
Lauffer, BE; Mintzer, R; Fong, R; Mukund, S; Tam, C; Zilberleyb, I; Flicke, B; Ritscher, A; Fedorowicz, G; Vallero, R; Ortwine, DF; Gunzner, J; Modrusan, Z; Neumann, L; Koth, CM; Lupardus, PJ; Kaminker, JS; Heise, CE; Steiner, P Histone deacetylase (HDAC) inhibitor kinetic rate constants correlate with cellular histone acetylation but not transcription and cell viability. J Biol Chem288:26926-43 (2013) [PubMed] Article