 Found 4 hits of Enzyme Inhibition Constant Data
Found 4 hits of Enzyme Inhibition Constant Data Target/Host
(Institution) | Ligand | Target/Host
Links | Ligand
Links | Trg + Lig
Links | Ki
nM | ΔG°
kJ/mole | IC50
nM | Kd
nM | EC50/IC50
nM | koff
s-1 | kon
M-1s-1 | pH | Temp
°C |
|---|
Heat shock protein HSP 90-alpha
(Homo sapiens (Human)) | BDBM20732
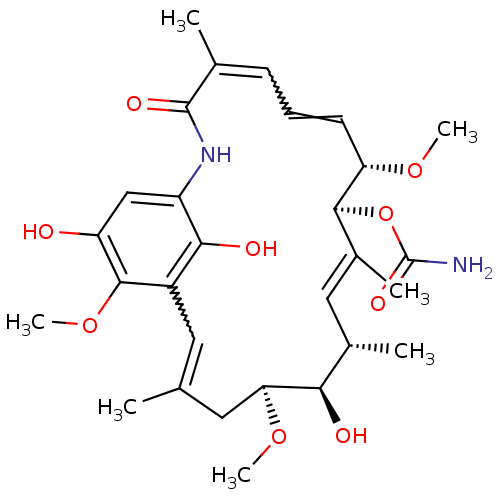
((4E,6Z,8S,9S,10E,12S,13R,14S,16R)-13-hydroxy-8,14,...)Show SMILES CO[C@H]1CC(C)=Cc2c(O)c(NC(=O)C(C)=CC=C[C@H](OC)[C@@H](OC(N)=O)\C(C)=C\[C@H](C)[C@H]1O)cc(O)c2OC |r,w:16.16,6.6,t:28| Show InChI InChI=1S/C29H40N2O9/c1-15-11-19-25(34)20(14-21(32)27(19)39-7)31-28(35)16(2)9-8-10-22(37-5)26(40-29(30)36)18(4)13-17(3)24(33)23(12-15)38-6/h8-11,13-14,17,22-24,26,32-34H,12H2,1-7H3,(H2,30,36)(H,31,35)/b10-8?,15-11?,16-9?,18-13+/t17-,22-,23-,24+,26-/m0/s1 | PDB
MMDB
NCI pathway
Reactome pathway
KEGG
UniProtKB/SwissProt
B.MOAD
DrugBank
antibodypedia
GoogleScholar
AffyNet 
| CHEMBL
MMDB
PC cid
PC sid
PDB
UniChem
Patents
Similars
| MMDB
PDB
Article
PubMed
| n/a | n/a | n/a | 1.22E+3 | n/a | n/a | n/a | n/a | n/a |
University College London
Curated by ChEMBL
| Assay Description
The compound was tested for binding affinity against yeast Heat shock protein HSP 90 |
J Med Chem 42: 260-6 (1999)
Article DOI: 10.1021/jm980403y
BindingDB Entry DOI: 10.7270/Q29G5NHQ |
More data for this
Ligand-Target Pair | 
3D Structure (crystal) |
Heat shock protein HSP 90-alpha
(Homo sapiens (Human)) | BDBM50347503
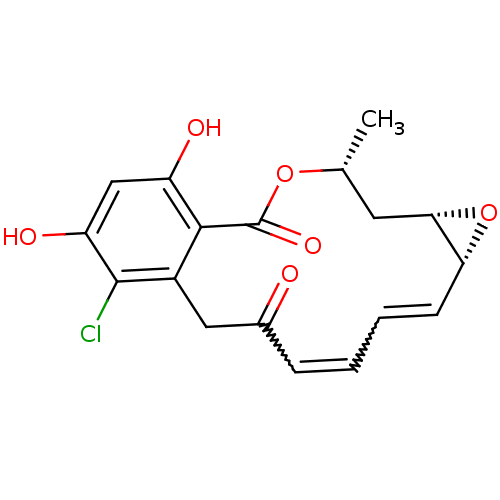
(RADICICOL)Show SMILES C[C@@H]1C[C@@H]2O[C@@H]2C=CC=CC(=O)Cc2c(Cl)c(O)cc(O)c2C(=O)O1 |r,w:9.10,7.8| Show InChI InChI=1S/C18H17ClO6/c1-9-6-15-14(25-15)5-3-2-4-10(20)7-11-16(18(23)24-9)12(21)8-13(22)17(11)19/h2-5,8-9,14-15,21-22H,6-7H2,1H3/t9-,14-,15+/m1/s1 | PDB
MMDB
NCI pathway
Reactome pathway
KEGG
UniProtKB/SwissProt
B.MOAD
DrugBank
antibodypedia
GoogleScholar
AffyNet 
| Purchase
PC cid
PC sid
UniChem
Patents
Similars
| Article
PubMed
| n/a | n/a | n/a | 19 | n/a | n/a | n/a | n/a | n/a |
University College London
Curated by ChEMBL
| Assay Description
The compound was tested for binding affinity against yeast Heat shock protein HSP 90 |
J Med Chem 42: 260-6 (1999)
Article DOI: 10.1021/jm980403y
BindingDB Entry DOI: 10.7270/Q29G5NHQ |
More data for this
Ligand-Target Pair | |
Heat shock protein HSP 90-alpha
(Homo sapiens (Human)) | BDBM50347503

(RADICICOL)Show SMILES C[C@@H]1C[C@@H]2O[C@@H]2C=CC=CC(=O)Cc2c(Cl)c(O)cc(O)c2C(=O)O1 |r,w:9.10,7.8| Show InChI InChI=1S/C18H17ClO6/c1-9-6-15-14(25-15)5-3-2-4-10(20)7-11-16(18(23)24-9)12(21)8-13(22)17(11)19/h2-5,8-9,14-15,21-22H,6-7H2,1H3/t9-,14-,15+/m1/s1 | PDB
MMDB
NCI pathway
Reactome pathway
KEGG
UniProtKB/SwissProt
B.MOAD
DrugBank
antibodypedia
GoogleScholar
AffyNet 
| Purchase
PC cid
PC sid
UniChem
Patents
Similars
| Article
PubMed
| n/a | n/a | n/a | 2.70 | n/a | n/a | n/a | n/a | n/a |
University College London
Curated by ChEMBL
| Assay Description
The compound was tested for binding affinity against isolated N-domain of Heat shock protein HSP 90 |
J Med Chem 42: 260-6 (1999)
Article DOI: 10.1021/jm980403y
BindingDB Entry DOI: 10.7270/Q29G5NHQ |
More data for this
Ligand-Target Pair | |
Heat shock protein HSP 90-alpha
(Homo sapiens (Human)) | BDBM20732

((4E,6Z,8S,9S,10E,12S,13R,14S,16R)-13-hydroxy-8,14,...)Show SMILES CO[C@H]1CC(C)=Cc2c(O)c(NC(=O)C(C)=CC=C[C@H](OC)[C@@H](OC(N)=O)\C(C)=C\[C@H](C)[C@H]1O)cc(O)c2OC |r,w:16.16,6.6,t:28| Show InChI InChI=1S/C29H40N2O9/c1-15-11-19-25(34)20(14-21(32)27(19)39-7)31-28(35)16(2)9-8-10-22(37-5)26(40-29(30)36)18(4)13-17(3)24(33)23(12-15)38-6/h8-11,13-14,17,22-24,26,32-34H,12H2,1-7H3,(H2,30,36)(H,31,35)/b10-8?,15-11?,16-9?,18-13+/t17-,22-,23-,24+,26-/m0/s1 | PDB
MMDB
NCI pathway
Reactome pathway
KEGG
UniProtKB/SwissProt
B.MOAD
DrugBank
antibodypedia
GoogleScholar
AffyNet 
| CHEMBL
MMDB
PC cid
PC sid
PDB
UniChem
Patents
Similars
| MMDB
PDB
Article
PubMed
| n/a | n/a | n/a | 775 | n/a | n/a | n/a | n/a | n/a |
University College London
Curated by ChEMBL
| Assay Description
The compound was tested for binding affinity against isolated N-domain of Heat shock protein HSP 90 |
J Med Chem 42: 260-6 (1999)
Article DOI: 10.1021/jm980403y
BindingDB Entry DOI: 10.7270/Q29G5NHQ |
More data for this
Ligand-Target Pair | 
3D Structure (crystal) |


 Search and Browse
Search and Browse
 Download
Download
 Enter Data
Enter Data