

 Search and Browse
Search and Browse
 Download
Download
 Enter Data
Enter Data
| Reaction Details | |||
|---|---|---|---|
 | Report a problem with these data | ||
| Cell Reactant: | HIV-1 Protease Mutant (V82F/I84V) | ||
| Syringe Reactant: | BDBM519 | ||
| Meas. Tech.: | Isothermal Titration Calorimetry | ||
| Entry Date: | 04/20/04 | ||
| ΔG°: | -43.5± (kJ/mole) | ||
| pH: | 5±n/a | ||
| Log10Kb: | n/a | ||
| Temperature: | 298.15±n/a (K) | ||
| ΔH° : | 14.6±0.627 (kJ/mole) | ||
| ΔHobs : | 14.4± (kJ/mole) | ||
| Ionic Strength: | n/a | ||
| not known | |||
| Protons Released: | 0.4 | ||
| ΔCp : | -1.7138±0.1672 (kJ/mole) | ||
| Stoich. Param.: | n/a | ||
| ΔS° : | 0.195± (kJ/mole-K) | ||
| Comments: | n/a | ||
| Citation |  Todd, MJ; Luque, I; Vel�zquez-Campoy, A; Freire, E Thermodynamic basis of resistance to HIV-1 protease inhibition: calorimetric analysis of the V82F/I84V active site resistant mutant. Biochemistry39:11876-83 (2000) [PubMed] Article Todd, MJ; Luque, I; Vel�zquez-Campoy, A; Freire, E Thermodynamic basis of resistance to HIV-1 protease inhibition: calorimetric analysis of the V82F/I84V active site resistant mutant. Biochemistry39:11876-83 (2000) [PubMed] Article | ||
| More Info.: | Get all data from this article , ITC RUN data , Solution Info , Data Fit Method , Instrument Info | ||
| HIV-1 Protease Mutant (V82F/I84V) | |||
| Source: | mutations at position 82 and 84 were introduced using an in vitro site-directed mutagenesis kit, mutant protease was expressed in BL21/DE3 | ||
| Purity: | 99% | ||
| Prep. Method: | HIV-1 protease was purified and refolded from E. coli inclusion bodies. | ||
| Name: | HIV-1 Protease Mutant (V82F/I84V) | ||
| Synonyms: | n/a | ||
| Type: | Protein Complex | ||
| Mol. Mass.: | n/a | ||
| Description: | n/a | ||
| Components: | This complex has 2 components. | ||
| Component 1 | |||
| Name: | HIV-1 Protease Mutant (V82F/I84V) chain A | ||
| Synonyms: | HIV-1 Protease Mutant (V82F/I84V) chain B | ||
| Type: | Enzyme Subunit | ||
| Mol. Mass.: | 10816.23 | ||
| Organism: | Human immunodeficiency virus type 1 | ||
| Description: | Using plasmid-encoded mutant protease (Q7K/L33I/L63I designed to remove three hypersensitive autolytic sites) for the stability of protease. | ||
| Residue: | 99 | ||
| Sequence: |
| ||
| Component 2 | |||
| Name: | HIV-1 Protease Mutant (V82F/I84V) chain A | ||
| Synonyms: | HIV-1 Protease Mutant (V82F/I84V) chain B | ||
| Type: | Enzyme Subunit | ||
| Mol. Mass.: | 10816.23 | ||
| Organism: | Human immunodeficiency virus type 1 | ||
| Description: | Using plasmid-encoded mutant protease (Q7K/L33I/L63I designed to remove three hypersensitive autolytic sites) for the stability of protease. | ||
| Residue: | 99 | ||
| Sequence: |
| ||
| BDBM519 | |||
| Source: | Purified from commercial capsules | ||
| Purity: | n/a | ||
| Prep. Method: | Further purified by HPLC using a semipreparative C-18 reversed-phase column developed with 0-100% acetonitrile in 0.05% TFA | ||
| Name | BDBM519 | ||
| Synonyms: | (2S)-N-[(2S,3R)-4-[(3S,4aS,8aS)-3-(tert-butylcarbamoyl)-decahydroisoquinolin-2-yl]-3-hydroxy-1-phenylbutan-2-yl]-2-(quinolin-2-ylformamido)butanediamide | CHEMBL114 | Fortovase | Invirase | Ro 31-8959 | SQV | Saquinavir | ||
| Type | Small organic molecule | ||
| Emp. Form. | C38H50N6O5 | ||
| Mol. Mass. | 670.8408 | ||
| SMILES | CC(C)(C)NC(=O)[C@@H]1C[C@@H]2CCCC[C@@H]2CN1C[C@@H](O)[C@H](Cc1ccccc1)NC(=O)[C@H](CC(N)=O)NC(=O)c1ccc2ccccc2n1 |@:16| | ||
| Structure | 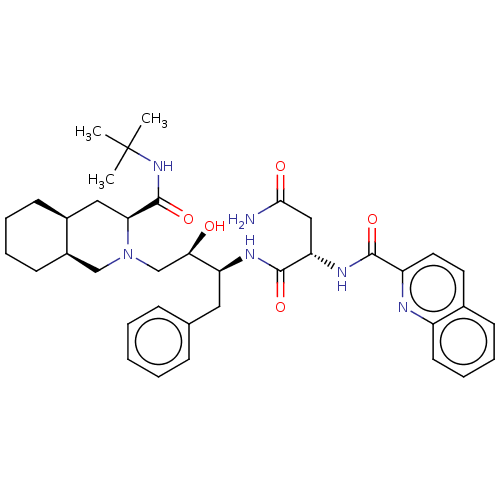
| ||
|
Home |
| |
Search |
| |
Deposit |
| |
SiteMap |
| |
About us |
| |
Email us |
| |
Info |
|
©2000 BindingDB. All rights reserved. |
|||||||||||||