

 Search and Browse
Search and Browse
 Download
Download
 Enter Data
Enter Data
| Reaction Details | |||
|---|---|---|---|
 | Report a problem with these data | ||
| Cell Reactant: | HIV-1 Protease | ||
| Syringe Reactant: | BDBM519 | ||
| Meas. Tech.: | Isothermal Titration Calorimetry | ||
| Entry Date: | 06/23/04 | ||
| ΔG°: | -12.8± (kcal/mole) | ||
| pH: | 5±n/a | ||
| Log10Kb: | 9.40± 7.80 | ||
| Temperature: | 298.15±n/a (K) | ||
| ΔH° : | 1.90± (kcal/mole) | ||
| ΔHobs : | 1.90± (kcal/mole) | ||
| Ionic Strength: | n/a | ||
| not known | |||
| Protons Released: | n/a | ||
| ΔCp : | n/a | ||
| Stoich. Param.: | n/a | ||
| ΔS° : | 0.0500± (kcal/mole-K) | ||
| Comments: | Adapted from Velazquez-Campoy, A., Kiso, Y., and Freire, E. (2001) Arch. Biochem. Biophys. 390, 169-175. | ||
| Citation |  Muzammil, S; Muzammil, S; Ross, P; Ross, P; Freire, E; Freire, E A major role for a set of non-active site mutations in the development of HIV-1 protease drug resistance. Biochemistry42:631-8 (2003) [PubMed] Article Muzammil, S; Muzammil, S; Ross, P; Ross, P; Freire, E; Freire, E A major role for a set of non-active site mutations in the development of HIV-1 protease drug resistance. Biochemistry42:631-8 (2003) [PubMed] Article | ||
| More Info.: | Get all data from this article , ITC RUN data , Solution Info , Data Fit Method , Instrument Info | ||
| HIV-1 Protease | |||
| Source: | Plasmid-encoded HIV-1 protease was expressed in E. coli cells. | ||
| Purity: | 99% | ||
| Prep. Method: | HIV-1 protease was purified and refolded from E. coli inclusion bodies. | ||
| Name: | Dimer of Gag-Pol polyprotein [489-587,Q496K] | ||
| Synonyms: | HIV-1 Protease | HIV-1 Protease (Q7K) | ||
| Type: | Protein Complex | ||
| Mol. Mass.: | n/a | ||
| Description: | n/a | ||
| Components: | This complex has 2 components. | ||
| Component 1 | |||
| Name: | Gag-Pol polyprotein [489-587,Q496K] | ||
| Synonyms: | HIV-1 Protease (Q7K) chain A | HIV-1 Protease (Q7K) chain B | HIV-1 Protease B Subtype chain A | HIV-1 Protease B Subtype chain B | HIV-1 Protease Mutant (Q7K) | HIV-1 Protease chain A | POL_HV1H2 | gag-pol | ||
| Type: | Enzyme Subunit | ||
| Mol. Mass.: | 10782.21 | ||
| Organism: | Human immunodeficiency virus type 1 | ||
| Description: | All of the protease constructs in this study are on the wild type containing the single Q7K mutation to prevent autocatalysis.This protein has enzymatic properties similar to that of the wild type protease. | ||
| Residue: | 99 | ||
| Sequence: |
| ||
| Component 2 | |||
| Name: | Gag-Pol polyprotein [489-587,Q496K] | ||
| Synonyms: | HIV-1 Protease (Q7K) chain A | HIV-1 Protease (Q7K) chain B | HIV-1 Protease B Subtype chain A | HIV-1 Protease B Subtype chain B | HIV-1 Protease Mutant (Q7K) | HIV-1 Protease chain A | POL_HV1H2 | gag-pol | ||
| Type: | Enzyme Subunit | ||
| Mol. Mass.: | 10782.21 | ||
| Organism: | Human immunodeficiency virus type 1 | ||
| Description: | All of the protease constructs in this study are on the wild type containing the single Q7K mutation to prevent autocatalysis.This protein has enzymatic properties similar to that of the wild type protease. | ||
| Residue: | 99 | ||
| Sequence: |
| ||
| BDBM519 | |||
| Source: | Purified from commercial capsules | ||
| Purity: | n/a | ||
| Prep. Method: | Further purified by HPLC using a semipreparative C-18 reversed-phase column developed with 0-100% acetonitrile in 0.05% TFA | ||
| Name | BDBM519 | ||
| Synonyms: | (2S)-N-[(2S,3R)-4-[(3S,4aS,8aS)-3-(tert-butylcarbamoyl)-decahydroisoquinolin-2-yl]-3-hydroxy-1-phenylbutan-2-yl]-2-(quinolin-2-ylformamido)butanediamide | CHEMBL114 | Fortovase | Invirase | Ro 31-8959 | SQV | Saquinavir | ||
| Type | Small organic molecule | ||
| Emp. Form. | C38H50N6O5 | ||
| Mol. Mass. | 670.8408 | ||
| SMILES | CC(C)(C)NC(=O)[C@@H]1C[C@@H]2CCCC[C@@H]2CN1C[C@@H](O)[C@H](Cc1ccccc1)NC(=O)[C@H](CC(N)=O)NC(=O)c1ccc2ccccc2n1 |@:16| | ||
| Structure | 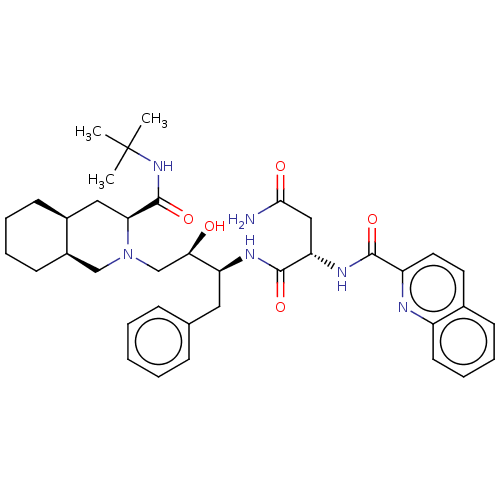
| ||
|
Home |
| |
Search |
| |
Deposit |
| |
SiteMap |
| |
About us |
| |
Email us |
| |
Info |
|
©2000 BindingDB. All rights reserved. |
|||||||||||||