| Reaction Details |
|---|
 | Report a problem with these data |
| Target | NADPH oxidase 1 |
|---|
| Ligand | BDBM32343 |
|---|
| Substrate/Competitor | n/a |
|---|
| Meas. Tech. | Late-stage luminescence-based cell-based dose response assay to identify inhibitors of NADPH oxidase 1 (NOX1): Purchased analogs |
|---|
| IC50 | >17000±n/a nM |
|---|
| Citation |  PubChem, PC Late-stage luminescence-based cell-based dose response assay to identify inhibitors of NADPH oxidase 1 (NOX1): Purchased analogs PubChem Bioassay(2010)[AID] PubChem, PC Late-stage luminescence-based cell-based dose response assay to identify inhibitors of NADPH oxidase 1 (NOX1): Purchased analogs PubChem Bioassay(2010)[AID] |
|---|
| More Info.: | Get all data from this article, Solution Info, Assay Method |
|---|
| |
| NADPH oxidase 1 |
|---|
| Name: | NADPH oxidase 1 |
|---|
| Synonyms: | MOX-1 | MOX1 | Mitogenic oxidase 1 | NADH/NADPH mitogenic oxidase subunit P65-MOX | NOH-1 | NOH1 | NOX-1 | NOX1 | NOX1_HUMAN |
|---|
| Type: | PROTEIN |
|---|
| Mol. Mass.: | 64890.44 |
|---|
| Organism: | Homo sapiens (Human) |
|---|
| Description: | ChEMBL_792877 |
|---|
| Residue: | 564 |
|---|
| Sequence: | MGNWVVNHWFSVLFLVVWLGLNVFLFVDAFLKYEKADKYYYTRKILGSTLACARASALCL
NFNSTLILLPVCRNLLSFLRGTCSFCSRTLRKQLDHNLTFHKLVAYMICLHTAIHIIAHL
FNFDCYSRSRQATDGSLASILSSLSHDEKKGGSWLNPIQSRNTTVEYVTFTSIAGLTGVI
MTIALILMVTSATEFIRRSYFEVFWYTHHLFIFYILGLGIHGIGGIVRGQTEESMNESHP
RKCAESFEMWDDRDSHCRRPKFEGHPPESWKWILAPVILYICERILRFYRSQQKVVITKV
VMHPSKVLELQMNKRGFSMEVGQYIFVNCPSISLLEWHPFTLTSAPEEDFFSIHIRAAGD
WTENLIRAFEQQYSPIPRIEVDGPFGTASEDVFQYEVAVLVGAGIGVTPFASILKSIWYK
FQCADHNLKTKKIYFYWICRETGAFSWFNNLLTSLEQEMEELGKVGFLNYRLFLTGWDSN
IVGHAALNFDKATDIVTGLKQKTSFGRPMWDNEFSTIATSHPKSVVGVFLCGPRTLAKSL
RKCCHRYSSLDPRKVQFYFNKENF
|
|
|
|---|
| BDBM32343 |
|---|
| n/a |
|---|
| Name | BDBM32343 |
|---|
| Synonyms: | (3,5-dichloro-4-pyridyl)-(4-pyridylmethyleneamino)amine | (3,5-dichloro-4-pyridyl)-[(E)-4-pyridylmethyleneamino]amine | 3,5-bis(chloranyl)-N-(pyridin-4-ylmethylideneamino)pyridin-4-amine | 3,5-dichloro-N-(pyridin-4-ylmethylideneamino)-4-pyridinamine | 3,5-dichloro-N-(pyridin-4-ylmethylideneamino)pyridin-4-amine | MLS000583434 | SMR000206620 | cid_4438738 | isonicotinaldehyde (3,5-dichloropyridin-4-yl)hydrazone |
|---|
| Type | Small organic molecule |
|---|
| Emp. Form. | C11H8Cl2N4 |
|---|
| Mol. Mass. | 267.114 |
|---|
| SMILES | Clc1cncc(Cl)c1N=NCc1ccncc1 |w:8.8| |
|---|
| Structure | 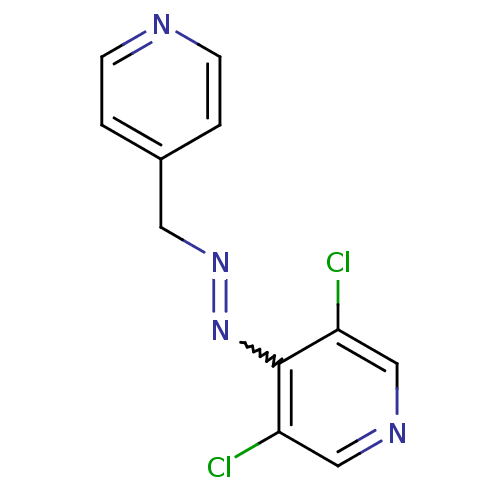
|
|---|


 Search and Browse
Search and Browse
 Download
Download
 Enter Data
Enter Data

 PubChem, PC Late-stage luminescence-based cell-based dose response assay to identify inhibitors of NADPH oxidase 1 (NOX1): Purchased analogs PubChem Bioassay(2010)[AID]
PubChem, PC Late-stage luminescence-based cell-based dose response assay to identify inhibitors of NADPH oxidase 1 (NOX1): Purchased analogs PubChem Bioassay(2010)[AID]