| Reaction Details |
|---|
 | Report a problem with these data |
| Target | Mu-type opioid receptor |
|---|
| Ligand | BDBM21864 |
|---|
| Substrate/Competitor | n/a |
|---|
| Meas. Tech. | ChEMBL_149639 (CHEMBL753634) |
|---|
| Ki | 34.4±n/a nM |
|---|
| Citation |  Metzger, TG; Paterlini, MG; Ferguson, DM; Portoghese, PS Investigation of the selectivity of oxymorphone- and naltrexone-derived ligands via site-directed mutagenesis of opioid receptors: exploring the"address" recognition locus. J Med Chem44:857-62 (2001) [PubMed] Metzger, TG; Paterlini, MG; Ferguson, DM; Portoghese, PS Investigation of the selectivity of oxymorphone- and naltrexone-derived ligands via site-directed mutagenesis of opioid receptors: exploring the"address" recognition locus. J Med Chem44:857-62 (2001) [PubMed] |
|---|
| More Info.: | Get all data from this article, Assay Method |
|---|
| |
| Mu-type opioid receptor |
|---|
| Name: | Mu-type opioid receptor |
|---|
| Synonyms: | MOR-1 | MUOR1 | Mu-type opioid receptor (MOR) | OPIATE Mu | OPRM_RAT | Opiate non-selective | Opioid receptor B | Oprm1 | Ror-b |
|---|
| Type: | G Protein-Coupled Receptor (GPCR) |
|---|
| Mol. Mass.: | 44503.11 |
|---|
| Organism: | Rattus norvegicus (rat) |
|---|
| Description: | Competition binding assays were carried out using membrane preparations from transfected HN9.10 cells that constitutively expressed the mu opioid receptor. |
|---|
| Residue: | 398 |
|---|
| Sequence: | MDSSTGPGNTSDCSDPLAQASCSPAPGSWLNLSHVDGNQSDPCGLNRTGLGGNDSLCPQT
GSPSMVTAITIMALYSIVCVVGLFGNFLVMYVIVRYTKMKTATNIYIFNLALADALATST
LPFQSVNYLMGTWPFGTILCKIVISIDYYNMFTSIFTLCTMSVDRYIAVCHPVKALDFRT
PRNAKIVNVCNWILSSAIGLPVMFMATTKYRQGSIDCTLTFSHPTWYWENLLKICVFIFA
FIMPVLIITVCYGLMILRLKSVRMLSGSKEKDRNLRRITRMVLVVVAVFIVCWTPIHIYV
IIKALITIPETTFQTVSWHFCIALGYTNSCLNPVLYAFLDENFKRCFREFCIPTSSTIEQ
QNSTRVRQNTREHPSTANTVDRTNHQLENLEAETAPLP
|
|
|
|---|
| BDBM21864 |
|---|
| n/a |
|---|
| Name | BDBM21864 |
|---|
| Synonyms: | (21R)-22-(cyclopropylmethyl)-14-oxa-11,22-diazaheptacyclo[13.9.1.0^{1,13}.0^{2,21}.0^{4,12}.0^{5,10}.0^{19,25}]pentacosa-4(12),5,7,9,15,17,19(25)-heptaene-2,16-diol | (8R)-7-(cyclopropylmethyl)-5,6,7,8,14,14b-hexahydro-4,8-methano[1]benzofuro[2,3-a]pyrido[4,3-b]carbazole-1,8a(9H)-diol | NALTRINDOLE | [3H]-naltrindole |
|---|
| Type | radiolabeled ligand |
|---|
| Emp. Form. | C26H26N2O3 |
|---|
| Mol. Mass. | 414.4962 |
|---|
| SMILES | [H][C@@]12Cc3ccc(O)c4OC5c6[nH]c7ccccc7c6CC1(O)C5(CCN2CC1CC1)c34 |THB:27:26:21:31.2.3| |
|---|
| Structure | 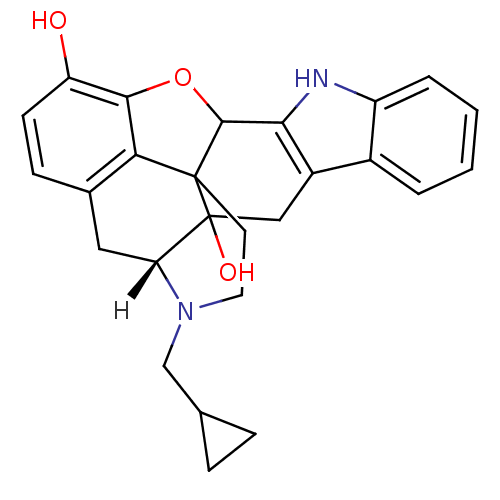
|
|---|


 Search and Browse
Search and Browse
 Download
Download
 Enter Data
Enter Data

 Metzger, TG; Paterlini, MG; Ferguson, DM; Portoghese, PS Investigation of the selectivity of oxymorphone- and naltrexone-derived ligands via site-directed mutagenesis of opioid receptors: exploring the"address" recognition locus. J Med Chem44:857-62 (2001) [PubMed]
Metzger, TG; Paterlini, MG; Ferguson, DM; Portoghese, PS Investigation of the selectivity of oxymorphone- and naltrexone-derived ligands via site-directed mutagenesis of opioid receptors: exploring the"address" recognition locus. J Med Chem44:857-62 (2001) [PubMed]