| Reaction Details |
|---|
 | Report a problem with these data |
| Target | UDP-3-O-acyl-N-acetylglucosamine deacetylase |
|---|
| Ligand | BDBM92264 |
|---|
| Substrate/Competitor | n/a |
|---|
| Meas. Tech. | Thermofluor |
|---|
| pH | 7±n/a |
|---|
| Temperature | 298.15±n/a K |
|---|
| Kd | >100000±n/a nM |
|---|
| Citation |  CSARdock, CD LPXC in CSAR_FULL_RELEASE_3JULY2012 CSAR1:0 (2012) CSARdock, CD LPXC in CSAR_FULL_RELEASE_3JULY2012 CSAR1:0 (2012) |
|---|
| More Info.: | Get all data from this article, Solution Info, Assay Method |
|---|
| |
| UDP-3-O-acyl-N-acetylglucosamine deacetylase |
|---|
| Name: | UDP-3-O-acyl-N-acetylglucosamine deacetylase |
|---|
| Synonyms: | LPXC_AQUAE | UDP-3-O-Acyl-GlcNAc Deacetylase (LpxC) | UDP-3-O-[3-hydroxymyristoyl] N-acetylglucosamine deacetylase | envA | lpxC | zinc deacetylase LpxC |
|---|
| Type: | Hydrolase |
|---|
| Mol. Mass.: | 32146.02 |
|---|
| Organism: | Aquifex aeolicus |
|---|
| Description: | The enzyme was stripped of all metal ions by dialysis against 1.0 mM EDTA in buffer at room temperature. The EDTA was then removed by extensive dialysis against EDTA-free buffer and the enzyme was reconstituted to a 1:1 Zn2+ to LpxC ratio by the addition of ZnSO4. |
|---|
| Residue: | 282 |
|---|
| Sequence: | MGLEKTVKEKLSFEGVGIHTGEYSKLIIHPEKEGTGIRFFKNGVYIPARHEFVVHTNHST
DLGFKGQRIKTVEHILSVLHLLEITNVTIEVIGNEIPILDGSGWEFYEAIRKNILNQNRE
IDYFVVEEPIIVEDEGRLIKAEPSDTLEVTYEGEFKNFLGRQKFTFVEGNEEEIVLARTF
CFDWEIEHIKKVGLGKGGSLKNTLVLGKDKVYNPEGLRYENEPVRHKVFDLIGDLYLLGS
PVKGKFYSFRGGHSLNVKLVKELAKKQKLTRDLPHLPSVQAL
|
|
|
|---|
| BDBM92264 |
|---|
| n/a |
|---|
| Name | BDBM92264 |
|---|
| Synonyms: | CS254 |
|---|
| Type | n/a |
|---|
| Emp. Form. | C24H24N4O5 |
|---|
| Mol. Mass. | 448.4712 |
|---|
| SMILES | C[C@@H](O)[C@H](NC(=O)c1ccc(cc1)C#Cc1ccccc1)C(=O)NCC1(C)NC(=O)NC1=O |
|---|
| Structure | 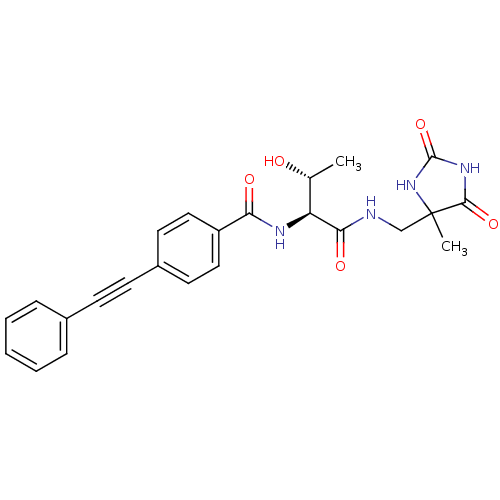
|
|---|


 Search and Browse
Search and Browse
 Download
Download
 Enter Data
Enter Data

 CSARdock, CD LPXC in CSAR_FULL_RELEASE_3JULY2012 CSAR1:0 (2012)
CSARdock, CD LPXC in CSAR_FULL_RELEASE_3JULY2012 CSAR1:0 (2012) 