| Reaction Details |
|---|
 | Report a problem with these data |
| Target | ATP-dependent molecular chaperone HSP82 |
|---|
| Ligand | BDBM265050 |
|---|
| Substrate/Competitor | n/a |
|---|
| Meas. Tech. | Malachite Green ATPase Assay |
|---|
| IC50 | >50000±n/a nM |
|---|
| Citation |  Drysdale, MJ; Dymock, BW; Finch, H; Webb, P; McDonald, E; James, KE; Cheung, KM; Matthews, TP Isoxazole compounds as inhibitors of heat shock proteins US Patent US11234987 Publication Date 2/1/2022 Drysdale, MJ; Dymock, BW; Finch, H; Webb, P; McDonald, E; James, KE; Cheung, KM; Matthews, TP Isoxazole compounds as inhibitors of heat shock proteins US Patent US11234987 Publication Date 2/1/2022 |
|---|
| More Info.: | Get all data from this article, Assay Method |
|---|
| |
| ATP-dependent molecular chaperone HSP82 |
|---|
| Name: | ATP-dependent molecular chaperone HSP82 |
|---|
| Synonyms: | 82 kDa heat shock protein | ATP-dependent molecular chaperone HSP82 | HSP82 | HSP82_YEAST | HSP90 | Heat Shock Protein 90 (Hsp90) | Heat shock protein Hsp90 heat-inducible isoform |
|---|
| Type: | Molecular Chaperone |
|---|
| Mol. Mass.: | 81369.08 |
|---|
| Organism: | Saccharomyces cerevisiae |
|---|
| Description: | Histidine-tagged yeast HSP90 was transformed into E. coli and purified (>90%) by metal affinity, gel filtration, and ion-exchange chromatography. |
|---|
| Residue: | 709 |
|---|
| Sequence: | MASETFEFQAEITQLMSLIINTVYSNKEIFLRELISNASDALDKIRYKSLSDPKQLETEP
DLFIRITPKPEQKVLEIRDSGIGMTKAELINNLGTIAKSGTKAFMEALSAGADVSMIGQF
GVGFYSLFLVADRVQVISKSNDDEQYIWESNAGGSFTVTLDEVNERIGRGTILRLFLKDD
QLEYLEEKRIKEVIKRHSEFVAYPIQLVVTKEVEKEVPIPEEEKKDEEKKDEEKKDEDDK
KPKLEEVDEEEEKKPKTKKVKEEVQEIEELNKTKPLWTRNPSDITQEEYNAFYKSISNDW
EDPLYVKHFSVEGQLEFRAILFIPKRAPFDLFESKKKKNNIKLYVRRVFITDEAEDLIPE
WLSFVKGVVDSEDLPLNLSREMLQQNKIMKVIRKNIVKKLIEAFNEIAEDSEQFEKFYSA
FSKNIKLGVHEDTQNRAALAKLLRYNSTKSVDELTSLTDYVTRMPEHQKNIYYITGESLK
AVEKSPFLDALKAKNFEVLFLTDPIDEYAFTQLKEFEGKTLVDITKDFELEETDEEKAER
EKEIKEYEPLTKALKEILGDQVEKVVVSYKLLDAPAAIRTGQFGWSANMERIMKAQALRD
SSMSSYMSSKKTFEISPKSPIIKELKKRVDEGGAQDKTVKDLTKLLYETALLTSGFSLDE
PTSFASRINRLISLGLNIDEDEETETAPEASTAAPVEEVPADTEMEEVD
|
|
|
|---|
| BDBM265050 |
|---|
| n/a |
|---|
| Name | BDBM265050 |
|---|
| Synonyms: | US10413550, Example 12 | US11234987, Example 12 | US9718793, 12 |
|---|
| Type | Small organic molecule |
|---|
| Emp. Form. | C22H18N2O4 |
|---|
| Mol. Mass. | 374.3893 |
|---|
| SMILES | COc1ccc(cc1)-c1c(C)noc1-c1cc(c(O)cc1O)-c1cccnc1 |
|---|
| Structure | 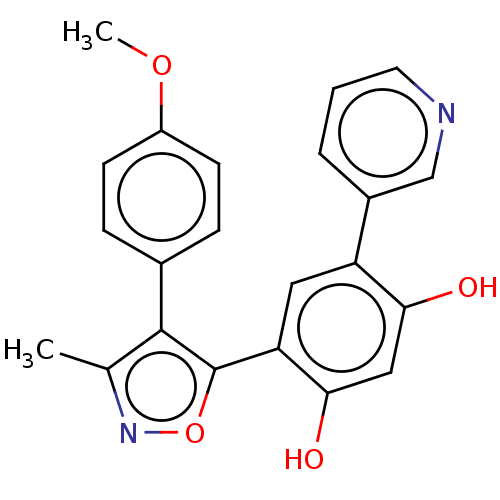
|
|---|


 Search and Browse
Search and Browse
 Download
Download
 Enter Data
Enter Data

 Drysdale, MJ; Dymock, BW; Finch, H; Webb, P; McDonald, E; James, KE; Cheung, KM; Matthews, TP Isoxazole compounds as inhibitors of heat shock proteins US Patent US11234987 Publication Date 2/1/2022
Drysdale, MJ; Dymock, BW; Finch, H; Webb, P; McDonald, E; James, KE; Cheung, KM; Matthews, TP Isoxazole compounds as inhibitors of heat shock proteins US Patent US11234987 Publication Date 2/1/2022