| Reaction Details |
|---|
 | Report a problem with these data |
| Target | Mitogen-activated protein kinase 14 |
|---|
| Ligand | BDBM14838 |
|---|
| Substrate/Competitor | BDBM14832 |
|---|
| Meas. Tech. | Fluorescence Exchange Curve Assay |
|---|
| pH | 7±n/a |
|---|
| Temperature | 296.15±n/a K |
|---|
| Kd | 6±n/a nM |
|---|
| Citation |  Kroe, RR; Regan, J; Proto, A; Peet, GW; Roy, T; Landro, LD; Fuschetto, NG; Pargellis, CA; Ingraham, RH Thermal denaturation: a method to rank slow binding, high-affinity P38alpha MAP kinase inhibitors. J Med Chem46:4669-75 (2003) [PubMed] Article Kroe, RR; Regan, J; Proto, A; Peet, GW; Roy, T; Landro, LD; Fuschetto, NG; Pargellis, CA; Ingraham, RH Thermal denaturation: a method to rank slow binding, high-affinity P38alpha MAP kinase inhibitors. J Med Chem46:4669-75 (2003) [PubMed] Article |
|---|
| More Info.: | Get all data from this article, Solution Info, Assay Method |
|---|
| |
| Mitogen-activated protein kinase 14 |
|---|
| Name: | Mitogen-activated protein kinase 14 |
|---|
| Synonyms: | Crk1 | Csbp1 | Csbp2 | MAP Kinase p38 alpha | MAP kinase p38 | MK14_MOUSE | Mapk14 | Mitogen-activated protein kinase 14 | Mitogen-activated protein kinase p38 alpha |
|---|
| Type: | Enzyme |
|---|
| Mol. Mass.: | 41281.22 |
|---|
| Organism: | Mus musculus (mouse) |
|---|
| Description: | The full-length open reading frame of murine p38 alpha was cloned and expressed in E. coli.. Soluble murine p38R was extracted from cell pellets and purified using ion-exchange chromatography. |
|---|
| Residue: | 360 |
|---|
| Sequence: | MSQERPTFYRQELNKTIWEVPERYQNLSPVGSGAYGSVCAAFDTKTGHRVAVKKLSRPFQ
SIIHAKRTYRELRLLKHMKHENVIGLLDVFTPARSLEEFNDVYLVTHLMGADLNNIVKCQ
KLTDDHVQFLIYQILRGLKYIHSADIIHRDLKPSNLAVNEDCELKILDFGLARHTDDEMT
GYVATRWYRAPEIMLNWMHYNQTVDIWSVGCIMAELLTGRTLFPGTDHIDQLKLILRLVG
TPGAELLKKISSESARNYIQSLAQMPKMNFANVFIGANPLAVDLLEKMLVLDSDKRITAA
QALAHAYFAQYHDPDDEPVADPYDQSFESRDLLIDEWKSLTYDEVISFVPPPLDQEEMES
|
|
|
|---|
| BDBM14838 |
|---|
| BDBM14832 |
|---|
| Name | BDBM14838 |
|---|
| Synonyms: | 1-(5-iso-Propyl-2-phenyl-2H-pyrazol-3-yl)-3-[4-(2-morpholin-4-yl-ethoxy)-naphthalen-1-yl]-urea | 1-{4-[2-(morpholin-4-yl)ethoxy]naphthalen-1-yl}-3-[1-phenyl-3-(propan-2-yl)-1H-pyrazol-5-yl]urea | BIRB-796 Analog 32 | CHEMBL105248 | diaryl urea compound 7 |
|---|
| Type | Small organic molecule |
|---|
| Emp. Form. | C29H33N5O3 |
|---|
| Mol. Mass. | 499.604 |
|---|
| SMILES | CC(C)c1cc(NC(=O)Nc2ccc(OCCN3CCOCC3)c3ccccc23)n(n1)-c1ccccc1 |
|---|
| Structure | 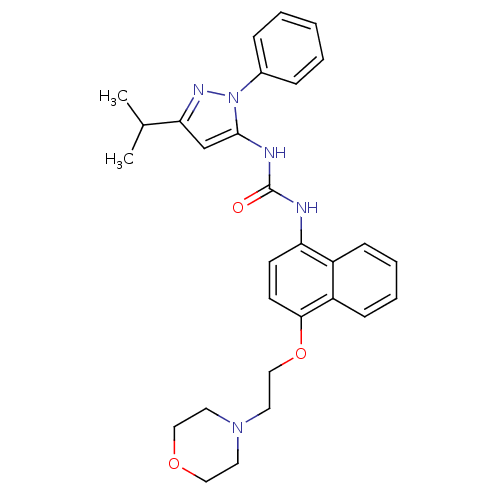
|
|---|


 Search and Browse
Search and Browse
 Download
Download
 Enter Data
Enter Data

 Kroe, RR; Regan, J; Proto, A; Peet, GW; Roy, T; Landro, LD; Fuschetto, NG; Pargellis, CA; Ingraham, RH Thermal denaturation: a method to rank slow binding, high-affinity P38alpha MAP kinase inhibitors. J Med Chem46:4669-75 (2003) [PubMed] Article
Kroe, RR; Regan, J; Proto, A; Peet, GW; Roy, T; Landro, LD; Fuschetto, NG; Pargellis, CA; Ingraham, RH Thermal denaturation: a method to rank slow binding, high-affinity P38alpha MAP kinase inhibitors. J Med Chem46:4669-75 (2003) [PubMed] Article