| Reaction Details |
|---|
 | Report a problem with these data |
| Target | G2/mitotic-specific cyclin-B1 |
|---|
| Ligand | BDBM43375 |
|---|
| Substrate/Competitor | n/a |
|---|
| Meas. Tech. | Counterscreen assay for inhibitors of Wee1 degradation: dose response cell-based assay to identify inhibitors of cyclin B degradation |
|---|
| EC50 | >1842±n/a nM |
|---|
| Citation |  PubChem, PC Counterscreen assay for inhibitors of Wee1 degradation: dose response cell-based assay to identify inhibitors of cyclin B degradation PubChem Bioassay(2008)[AID] PubChem, PC Counterscreen assay for inhibitors of Wee1 degradation: dose response cell-based assay to identify inhibitors of cyclin B degradation PubChem Bioassay(2008)[AID] |
|---|
| More Info.: | Get all data from this article, Solution Info, Assay Method |
|---|
| |
| G2/mitotic-specific cyclin-B1 |
|---|
| Name: | G2/mitotic-specific cyclin-B1 |
|---|
| Synonyms: | CCNB | CCNB1 | CCNB1_HUMAN |
|---|
| Type: | Enzyme Subunit |
|---|
| Mol. Mass.: | 48340.95 |
|---|
| Organism: | Homo sapiens (Human) |
|---|
| Description: | P14635 |
|---|
| Residue: | 433 |
|---|
| Sequence: | MALRVTRNSKINAENKAKINMAGAKRVPTAPAATSKPGLRPRTALGDIGNKVSEQLQAKM
PMKKEAKPSATGKVIDKKLPKPLEKVPMLVPVPVSEPVPEPEPEPEPEPVKEEKLSPEPI
LVDTASPSPMETSGCAPAEEDLCQAFSDVILAVNDVDAEDGADPNLCSEYVKDIYAYLRQ
LEEEQAVRPKYLLGREVTGNMRAILIDWLVQVQMKFRLLQETMYMTVSIIDRFMQNNCVP
KKMLQLVGVTAMFIASKYEEMYPPEIGDFAFVTDNTYTKHQIRQMEMKILRALNFGLGRP
LPLHFLRRASKIGEVDVEQHTLAKYLMELTMLDYDMVHFPPSQIAAGAFCLALKILDNGE
WTPTLQHYLSYTEESLLPVMQHLAKNVVMVNQGLTKHMTVKNKYATSKHAKISTLPQLNS
ALVQDLAKAVAKV
|
|
|
|---|
| BDBM43375 |
|---|
| n/a |
|---|
| Name | BDBM43375 |
|---|
| Synonyms: | (2E)-2-(1-cyano-2,2,2-trifluoro-ethylidene)-5-phenyl-1,3-oxathiole-4-carboxylic acid ethyl ester | (2E)-2-(1-cyano-2,2,2-trifluoroethylidene)-5-phenyl-1,3-oxathiole-4-carboxylic acid ethyl ester | 2-[1-Cyano-2,2,2-trifluoro-eth-(E)-ylidene]-5-phenyl-[1,3]oxathiole-4-carboxylic acid ethyl ester | MLS000556838 | SMR000147955 | cid_1883255 | ethyl (2E)-2-(1-cyano-2,2,2-trifluoroethylidene)-5-phenyl-1,3-oxathiole-4-carboxylate | ethyl (2E)-2-[1-cyano-2,2,2-tris(fluoranyl)ethylidene]-5-phenyl-1,3-oxathiole-4-carboxylate |
|---|
| Type | Small organic molecule |
|---|
| Emp. Form. | C15H10F3NO3S |
|---|
| Mol. Mass. | 341.305 |
|---|
| SMILES | CCOC(=O)C1=C(O\C(S1)=C(\C#N)C(F)(F)F)c1ccccc1 |t:5| |
|---|
| Structure | 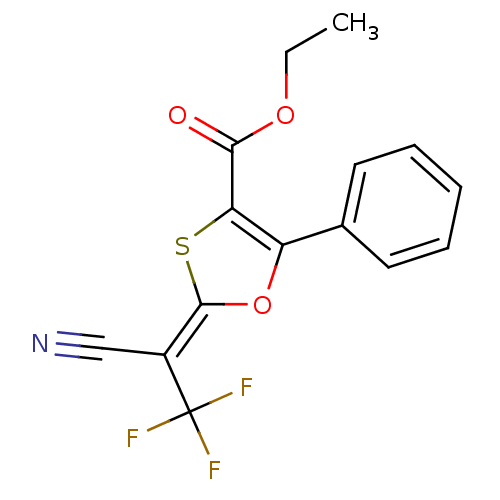
|
|---|


 Search and Browse
Search and Browse
 Download
Download
 Enter Data
Enter Data

 PubChem, PC Counterscreen assay for inhibitors of Wee1 degradation: dose response cell-based assay to identify inhibitors of cyclin B degradation PubChem Bioassay(2008)[AID]
PubChem, PC Counterscreen assay for inhibitors of Wee1 degradation: dose response cell-based assay to identify inhibitors of cyclin B degradation PubChem Bioassay(2008)[AID]