| Reaction Details |
|---|
 | Report a problem with these data |
| Target | Voltage-dependent T-type calcium channel subunit alpha-1H |
|---|
| Ligand | BDBM74444 |
|---|
| Substrate/Competitor | n/a |
|---|
| Meas. Tech. | Inhibitors of T-Type Calcium Channel |
|---|
| EC50 | 1650±0 nM |
|---|
| Citation |  PubChem, PC Inhibitors of T-Type Calcium Channel PubChem Bioassay(2010)[AID] PubChem, PC Inhibitors of T-Type Calcium Channel PubChem Bioassay(2010)[AID] |
|---|
| More Info.: | Get all data from this article, Solution Info, Assay Method |
|---|
| |
| Voltage-dependent T-type calcium channel subunit alpha-1H |
|---|
| Name: | Voltage-dependent T-type calcium channel subunit alpha-1H |
|---|
| Synonyms: | CAC1H_HUMAN | CACNA1H | Low-voltage-activated calcium channel alpha1 3.2 subunit | Voltage-gated T-type calcium channel | Voltage-gated calcium channel | Voltage-gated calcium channel subunit alpha Cav3.2 | voltage-dependent T-type calcium channel subunit alpha-1H isoform a |
|---|
| Type: | Enzyme Catalytic Domain |
|---|
| Mol. Mass.: | 259187.68 |
|---|
| Organism: | Homo sapiens (Human) |
|---|
| Description: | Calcium channel (Type T,L) 0 0::O95180 |
|---|
| Residue: | 2353 |
|---|
| Sequence: | MTEGARAADEVRVPLGAPPPGPAALVGASPESPGAPGREAERGSELGVSPSESPAAERGA
ELGADEEQRVPYPALAATVFFCLGQTTRPRSWCLRLVCNPWFEHVSMLVIMLNCVTLGMF
RPCEDVECGSERCNILEAFDAFIFAFFAVEMVIKMVALGLFGQKCYLGDTWNRLDFFIVV
AGMMEYSLDGHNVSLSAIRTVRVLRPLRAINRVPSMRILVTLLLDTLPMLGNVLLLCFFV
FFIFGIVGVQLWAGLLRNRCFLDSAFVRNNNLTFLRPYYQTEEGEENPFICSSRRDNGMQ
KCSHIPGRRELRMPCTLGWEAYTQPQAEGVGAARNACINWNQYYNVCRSGDSNPHNGAIN
FDNIGYAWIAIFQVITLEGWVDIMYYVMDAHSFYNFIYFILLIIVGSFFMINLCLVVIAT
QFSETKQRESQLMREQRARHLSNDSTLASFSEPGSCYEELLKYVGHIFRKVKRRSLRLYA
RWQSRWRKKVDPSAVQGQGPGHRQRRAGRHTASVHHLVYHHHHHHHHHYHFSHGSPRRPG
PEPGACDTRLVRAGAPPSPPSPGRGPPDAESVHSIYHADCHIEGPQERARVAHAAATAAA
SLRLATGLGTMNYPTILPSGVGSGKGSTSPGPKGKWAGGPPGTGGHGPLSLNSPDPYEKI
PHVVGEHGLGQAPGHLSGLSVPCPLPSPPAGTLTCELKSCPYCTRALEDPEGELSGSESG
DSDGRGVYEFTQDVRHGDRWDPTRPPRATDTPGPGPGSPQRRAQQRAAPGEPGWMGRLWV
TFSGKLRRIVDSKYFSRGIMMAILVNTLSMGVEYHEQPEELTNALEISNIVFTSMFALEM
LLKLLACGPLGYIRNPYNIFDGIIVVISVWEIVGQADGGLSVLRTFRLLRVLKLVRFLPA
LRRQLVVLVKTMDNVATFCTLLMLFIFIFSILGMHLFGCKFSLKTDTGDTVPDRKNFDSL
LWAIVTVFQILTQEDWNVVLYNGMASTSSWAALYFVALMTFGNYVLFNLLVAILVEGFQA
EGDANRSDTDEDKTSVHFEEDFHKLRELQTTELKMCSLAVTPNGHLEGRGSLSPPLIMCT
AATPMPTPKSSPFLDAAPSLPDSRRGSSSSGDPPLGDQKPPASLRSSPCAPWGPSGAWSS
RRSSWSSLGRAPSLKRRGQCGERESLLSGEGKGSTDDEAEDGRAAPGPRATPLRRAESLD
PRPLRPAALPPTKCRDRDGQVVALPSDFFLRIDSHREDAAELDDDSEDSCCLRLHKVLEP
YKPQWCRSREAWALYLFSPQNRFRVSCQKVITHKMFDHVVLVFIFLNCVTIALERPDIDP
GSTERVFLSVSNYIFTAIFVAEMMVKVVALGLLSGEHAYLQSSWNLLDGLLVLVSLVDIV
VAMASAGGAKILGVLRVLRLLRTLRPLRVISRAPGLKLVVETLISSLRPIGNIVLICCAF
FIIFGILGVQLFKGKFYYCEGPDTRNISTKAQCRAAHYRWVRRKYNFDNLGQALMSLFVL
SSKDGWVNIMYDGLDAVGVDQQPVQNHNPWMLLYFISFLLIVSFFVLNMFVGVVVENFHK
CRQHQEAEEARRREEKRLRRLERRRRSTFPSPEAQRRPYYADYSPTRRSIHSLCTSHYLD
LFITFIICVNVITMSMEHYNQPKSLDEALKYCNYVFTIVFVFEAALKLVAFGFRRFFKDR
WNQLDLAIVLLSLMGITLEEIEMSAALPINPTIIRIMRVLRIARVLKLLKMATGMRALLD
TVVQALPQVGNLGLLFMLLFFIYAALGVELFGRLECSEDNPCEGLSRHATFSNFGMAFLT
LFRVSTGDNWNGIMKDTLRECSREDKHCLSYLPALSPVYFVTFVLVAQFVLVNVVVAVLM
KHLEESNKEAREDAELDAEIELEMAQGPGSARRVDADRPPLPQESPGARDAPNLVARKVS
VSRMLSLPNDSYMFRPVVPASAPHPRPLQEVEMETYGAGTPLGSVASVHSPPAESCASLQ
IPLAVSSPARSGEPLHALSPRGTARSPSLSRLLCRQEAVHTDSLEGKIDSPRDTLDPAEP
GEKTPVRPVTQGGSLQSPPRSPRPASVRTRKHTFGQRCVSSRPAAPGGEEAEASDPADEE
VSHITSSACPWQPTAEPHGPEASPVAGGERDLRRLYSVDAQGFLDKPGRADEQWRPSAEL
GSGEPGEAKAWGPEAEPALGARRKKKMSPPCISVEPPAEDEGSARPSAAEGGSTTLRRRT
PSCEATPHRDSLEPTEGSGAGGDPAAKGERWGQASCRAEHLTVPSFAFEPLDLGVPSGDP
FLDGSHSVTPESRASSSGAIVPLEPPESEPPMPVGDPPEKRRGLYLTVPQCPLEKPGSPS
ATPAPGGGADDPV
|
|
|
|---|
| BDBM74444 |
|---|
| n/a |
|---|
| Name | BDBM74444 |
|---|
| Synonyms: | 4-Isopropyl-4H-furo[3,2-b]pyrrole-5-carboxylic acid [(benzo[1,3]dioxol-5-ylmethyl)-carbamoyl]-methyl ester | 4-isopropylfuro[3,2-b]pyrrole-5-carboxylic acid [2-keto-2-(piperonylamino)ethyl] ester | 4-propan-2-yl-5-furo[3,2-b]pyrrolecarboxylic acid [2-(1,3-benzodioxol-5-ylmethylamino)-2-oxoethyl] ester | MLS000035170 | SMR000008063 | [2-(1,3-benzodioxol-5-ylmethylamino)-2-oxidanylidene-ethyl] 4-propan-2-ylfuro[3,2-b]pyrrole-5-carboxylate | [2-(1,3-benzodioxol-5-ylmethylamino)-2-oxoethyl] 4-propan-2-ylfuro[3,2-b]pyrrole-5-carboxylate | cid_648085 |
|---|
| Type | Small organic molecule |
|---|
| Emp. Form. | C20H20N2O6 |
|---|
| Mol. Mass. | 384.3826 |
|---|
| SMILES | CC(C)n1c(cc2occc12)C(=O)OCC(=O)NCc1ccc2OCOc2c1 |
|---|
| Structure | 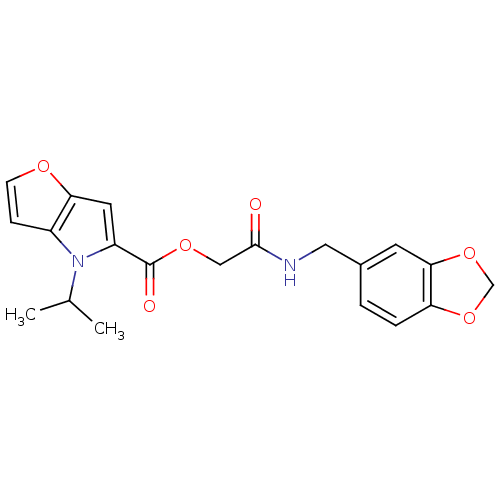
|
|---|


 Search and Browse
Search and Browse
 Download
Download
 Enter Data
Enter Data

 PubChem, PC Inhibitors of T-Type Calcium Channel PubChem Bioassay(2010)[AID]
PubChem, PC Inhibitors of T-Type Calcium Channel PubChem Bioassay(2010)[AID]