| Reaction Details |
|---|
 | Report a problem with these data |
| Target | Caspase-9 |
|---|
| Ligand | BDBM61198 |
|---|
| Substrate/Competitor | n/a |
|---|
| Meas. Tech. | Dose response confirmation of uHTS hits for Apaf-1 using a LZ-Caspase-9/Caspase-3 Fluorescent Selectivity assay |
|---|
| IC50 | 2630±n/a nM |
|---|
| Citation |  PubChem, PC Dose response confirmation of uHTS hits for Apaf-1 using a LZ-Caspase-9/Caspase-3 Fluorescent Selectivity assay PubChem Bioassay(2011)[AID] PubChem, PC Dose response confirmation of uHTS hits for Apaf-1 using a LZ-Caspase-9/Caspase-3 Fluorescent Selectivity assay PubChem Bioassay(2011)[AID] |
|---|
| More Info.: | Get all data from this article, Assay Method |
|---|
| |
| Caspase-9 |
|---|
| Name: | Caspase-9 |
|---|
| Synonyms: | Apoptotic protease-activating factor 1/Caspase-3/Caspase-9/Cytochrome c | Apoptotic protease-activating factor 3 | CASP-9 | CASP9 | CASP9_HUMAN | Caspase 9 | ICE-like apoptotic protease 6 | MCH6 | caspase-9 isoform alpha preproprotein |
|---|
| Type: | Enzyme |
|---|
| Mol. Mass.: | 46277.29 |
|---|
| Organism: | Homo sapiens (Human) |
|---|
| Description: | n/a |
|---|
| Residue: | 416 |
|---|
| Sequence: | MDEADRRLLRRCRLRLVEELQVDQLWDALLSRELFRPHMIEDIQRAGSGSRRDQARQLII
DLETRGSQALPLFISCLEDTGQDMLASFLRTNRQAAKLSKPTLENLTPVVLRPEIRKPEV
LRPETPRPVDIGSGGFGDVGALESLRGNADLAYILSMEPCGHCLIINNVNFCRESGLRTR
TGSNIDCEKLRRRFSSLHFMVEVKGDLTAKKMVLALLELAQQDHGALDCCVVVILSHGCQ
ASHLQFPGAVYGTDGCPVSVEKIVNIFNGTSCPSLGGKPKLFFIQACGGEQKDHGFEVAS
TSPEDESPGSNPEPDATPFQEGLRTFDQLDAISSLPTPSDIFVSYSTFPGFVSWRDPKSG
SWYVETLDDIFEQWAHSEDLQSLLLRVANAVSVKGIYKQMPGCFNFLRKKLFFKTS
|
|
|
|---|
| BDBM61198 |
|---|
| n/a |
|---|
| Name | BDBM61198 |
|---|
| Synonyms: | 2-methoxybenzaldehyde [2,6-bis(dimethylamino)pyrimidin-4-yl]hydrazone | 6-N-[(2-methoxyphenyl)methylideneamino]-2-N,2-N,4-N,4-N-tetramethylpyrimidine-2,4,6-triamine | MLS000720211 | N6-[(2-methoxyphenyl)methylideneamino]-N2,N2,N4,N4-tetramethyl-pyrimidine-2,4,6-triamine | N6-[(2-methoxyphenyl)methylideneamino]-N2,N2,N4,N4-tetramethylpyrimidine-2,4,6-triamine | SMR000304740 | [2-(dimethylamino)-6-(N'-o-anisylidenehydrazino)pyrimidin-4-yl]-dimethyl-amine | cid_4035805 |
|---|
| Type | Small organic molecule |
|---|
| Emp. Form. | C16H22N6O |
|---|
| Mol. Mass. | 314.3855 |
|---|
| SMILES | COc1ccccc1CN=Nc1cc(nc(n1)N(C)C)N(C)C |w:10.11| |
|---|
| Structure | 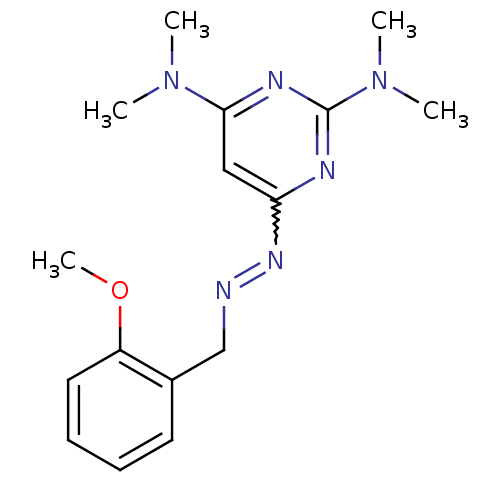
|
|---|


 Search and Browse
Search and Browse
 Download
Download
 Enter Data
Enter Data

 PubChem, PC Dose response confirmation of uHTS hits for Apaf-1 using a LZ-Caspase-9/Caspase-3 Fluorescent Selectivity assay PubChem Bioassay(2011)[AID]
PubChem, PC Dose response confirmation of uHTS hits for Apaf-1 using a LZ-Caspase-9/Caspase-3 Fluorescent Selectivity assay PubChem Bioassay(2011)[AID]