| Reaction Details |
|---|
 | Report a problem with these data |
| Target | Voltage-dependent L-type calcium channel subunit alpha-1C |
|---|
| Ligand | BDBM50029426 |
|---|
| Substrate/Competitor | n/a |
|---|
| Meas. Tech. | ChEMBL_92797 (CHEMBL703975) |
|---|
| Ki | 8.3±n/a nM |
|---|
| Citation |  Campiani, G; Garofalo, A; Fiorini, I; Botta, M; Nacci, V; Tafi, A; Chiarini, A; Budriesi, R; Bruni, G; Romeo, MR Pyrrolo[2,1-c][1,4]benzothiazines: synthesis, structure-activity relationships, molecular modeling studies, and cardiovascular activity. J Med Chem38:4393-410 (1995) [PubMed] Campiani, G; Garofalo, A; Fiorini, I; Botta, M; Nacci, V; Tafi, A; Chiarini, A; Budriesi, R; Bruni, G; Romeo, MR Pyrrolo[2,1-c][1,4]benzothiazines: synthesis, structure-activity relationships, molecular modeling studies, and cardiovascular activity. J Med Chem38:4393-410 (1995) [PubMed] |
|---|
| More Info.: | Get all data from this article, Assay Method |
|---|
| |
| Voltage-dependent L-type calcium channel subunit alpha-1C |
|---|
| Name: | Voltage-dependent L-type calcium channel subunit alpha-1C |
|---|
| Synonyms: | CAC1C_RAT | Cach2 | Cacn2 | Cacna1c | Cacnl1a1 | Calcium channel | Calcium channel (Type L) | Calcium channel diltiazem | Calcium channel, L type, alpha-1 polypeptide, isoform 1, cardiac muscle | Cchl1a1 | RBC | Rat brain class C | Voltage-dependent L-type calcium channel subunit alpha-1C | Voltage-gated L-type calcium channel | Voltage-gated L-type calcium channel alpha-1C subunit | Voltage-gated calcium channel subunit alpha Cav1.2 |
|---|
| Type: | Enzyme Catalytic Domain |
|---|
| Mol. Mass.: | 243492.26 |
|---|
| Organism: | RAT |
|---|
| Description: | Calcium channel 0 RAT::P22002 |
|---|
| Residue: | 2169 |
|---|
| Sequence: | MIRAFAQPSTPPYQPLSSCLSEDTERKFKGKVVHEAQLNCFYISPGGSNYGSPRPAHANM
NANAAAGLAPEHIPTPGAALSWLAAIDAARQAKLMGSAGNATISTVSSTQRKRQQYGKPK
KQGGTTATRPPRALLCLTLKNPIRRACISIVEWKPFEIIILLTIFANCVALAIYIPFPED
DSNATNSNLERVEYLFLIIFTVEAFLKVIAYGLLFHPNAYLRNGWNLLDFIIVVVGLFSA
ILEQATKADGANALGGKGAGFDVKALRAFRVLRPLRLVSGVPSLQVVLNSIIKAMVPLLH
IALLVLFVIIIYAIIGLELFMGKMHKTCYNQEGIIDVPAEEDPSPCALETGHGRQCQNGT
VCKPGWDGPKHGITNFDNFAFAMLTVFQCITMEGWTDVLYWMQDAMGYELPWVYFVSLVI
FGSFFVLNLVLGVLSGEFSKEREKAKARGDFQKLREKQQLEEDLKGYLDWITQAEDIDPE
NEDEGMDEDKPRNMSMPTSETESVNTENVAGGDIEGENCGARLAHRISKSKFSRYWRRWN
RFCRRKCRAAVKSNVFYWLVIFLVFLNTLTIASEHYNQPHWLTEVQDTANKALLALFTAE
MLLKMYSLGLQAYFVSLFNRFDCFIVCGGILETILVETKIMSPLGISCWRCVRLLRIFKI
TRYWNSLSNLVASLLNSLRSIASLLLLLFLFIIIFSLLGMQLFGGKFNFDEMQTRRSTFD
NFPQSLLTVFQILTGEDWNSVMYDGIMAYGGPSFPGMLVCIYFIILFISPNYILLNLFLA
IAVDNLADAESLTSAQKEEEEEKERKKLARTASPEKKQEVMEKPAVEESKEEKIELKSIT
ADGESPPTTKINMDDLQPSENEDKSPHSNPDTAGEEDEEEPEMPVGPRPRPLSELHLKEK
AVPMPEASAFFIFSPNNRFRLQCHRIVNDTIFTNLILFFILLSSISLAAEDPVQHTSFRN
HILFYFDIVFTTIFTIEIALKMTAYGAFLHKGSFCRNYFNILDLLVVSVSLISFGIQSSA
INVVKILRVLRVLRPLRINRAKGLKHVVQCVFVAIRTIGNIVIVTTLLQFMFACIGVQLF
KGKLYTCSDSSKQTEAESKGNYITYKTGEVDHPIIQPRSWENSKFDFDNVLAAMMALFTV
STFEGWPELLYRSIDSHTEDKGPIYNYRVEISIFFIIYIIIIAFFMMNIFVGFVIVTFQE
QGEQEYKNCELDKNQRQCVEYALKARPLPRYIPKNQHQYKVWYVVNSTYFEYLMFVLILL
NTICLAMQHYGQSCLFKIAMNILNMLFTGLFTVEMILKLIAFKPKHYFCDAWNTFDALIV
VGSIVDIAITEVHPAEHTQCSPSMSAEENSRISITFFRLFRVMRLVKLLSRGEGIRTLLW
TFIKSFQALPYVALLIVMLFFIYAVIGMQVFGKIALNDTTEINRNNNFQTFPQAVLLLFR
CATGEAWQDIMLACMPGKKCAPESEPSNSTEGETPCGSSFAVFYFISFYMLCAFLIINLF
VAVIMDNFDYLTRDWSILGPHHLDEFKRIWAEYDPEAKGRIKHLDVVTLLRRIQPPLGFG
KLCPHRVACKRLVSMNMPLNSDGTVMFNATLFALVRTALRIKTEGNLEQANEELRAIIKK
IWKRTSMKLLDQVVPPAGDDEVTVGKFYATFLIQEYFRKFKKRKEQGLVGKPSQRNALSL
QAGLRTLHDIGPEIRRAISGDLTAEEELDKAMKEAVSAASEDDIFRRAGGLFGNHVSYYQ
SDSRSNFPQTFATQRPLHINKTGNNQADTESPSHEKLVDSTFTPSSYSSTGSNANINNAN
NTALGRFPHPAGYSSTVSTVEGHGPPLSPAVRVQEAAWKLSSKRCHSRESQGATVSQDMF
PDETRSSVRLSEEVEYCSEPSLLSTDILSYQDDENRQLTCLEEDKREIQPCPKRSFLRSA
SLGRRASFHLECLKRQKDQGGDISQKTALPLHLVHHQALAVAGLSPLLQRSHSPSTFPRP
RPTPPVTPGSRGRPLQPIPTLRLEGAESSEKLNSSFPSIHCSSWSEETTACSGGSSMARR
ARPVSLTVPSQAGAPGRQFHGSASSLVEAVLISEGLGQFAQDPKFIEVTTQELADACDMT
IEEMENAADNILSGGAQQSPNGTLLPFVNCRDPGQDRAVVPEDESCVYALGRGRSEEALP
DSRSYVSNL
|
|
|
|---|
| BDBM50029426 |
|---|
| n/a |
|---|
| Name | BDBM50029426 |
|---|
| Synonyms: | CHEMBL24675 | [2-(3,4-Dimethoxy-phenyl)-ethyl]-[4-(4-methoxy-phenyl)-4H-benzo[b]pyrrolo[1,2-d][1,4]thiazin-1-ylmethyl]-methyl-amine |
|---|
| Type | Small organic molecule |
|---|
| Emp. Form. | C30H32N2O3S |
|---|
| Mol. Mass. | 500.652 |
|---|
| SMILES | COc1ccc(cc1)C1Sc2ccccc2-n2c(CN(C)CCc3ccc(OC)c(OC)c3)ccc12 |
|---|
| Structure | 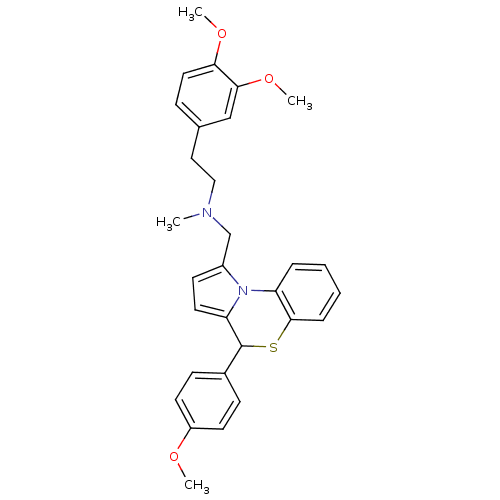
|
|---|


 Search and Browse
Search and Browse
 Download
Download
 Enter Data
Enter Data

 Campiani, G; Garofalo, A; Fiorini, I; Botta, M; Nacci, V; Tafi, A; Chiarini, A; Budriesi, R; Bruni, G; Romeo, MR Pyrrolo[2,1-c][1,4]benzothiazines: synthesis, structure-activity relationships, molecular modeling studies, and cardiovascular activity. J Med Chem38:4393-410 (1995) [PubMed]
Campiani, G; Garofalo, A; Fiorini, I; Botta, M; Nacci, V; Tafi, A; Chiarini, A; Budriesi, R; Bruni, G; Romeo, MR Pyrrolo[2,1-c][1,4]benzothiazines: synthesis, structure-activity relationships, molecular modeling studies, and cardiovascular activity. J Med Chem38:4393-410 (1995) [PubMed]