| Reaction Details |
|---|
 | Report a problem with these data |
| Target | Muscarinic acetylcholine receptor DM1 |
|---|
| Ligand | BDBM50064176 |
|---|
| Substrate/Competitor | n/a |
|---|
| Meas. Tech. | ChEMBL_906757 (CHEMBL3053219) |
|---|
| Ki | 3860±n/a nM |
|---|
| Citation |  Honda, H; Tomizawa, M; Casida, JE Insect muscarinic acetylcholine receptor: pharmacological and toxicological profiles of antagonists and agonists. J Agric Food Chem55:2276-81 (2007) [PubMed] Article Honda, H; Tomizawa, M; Casida, JE Insect muscarinic acetylcholine receptor: pharmacological and toxicological profiles of antagonists and agonists. J Agric Food Chem55:2276-81 (2007) [PubMed] Article |
|---|
| More Info.: | Get all data from this article, Assay Method |
|---|
| |
| Muscarinic acetylcholine receptor DM1 |
|---|
| Name: | Muscarinic acetylcholine receptor DM1 |
|---|
| Synonyms: | ACM1_DROME | AcrC | Muscarinic acetylcholine receptor DM1 | mAChR-A | mAcR-60C | mAcR-60C |
|---|
| Type: | PROTEIN |
|---|
| Mol. Mass.: | 86647.10 |
|---|
| Organism: | Drosophila melanogaster |
|---|
| Description: | ChEMBL_106611 |
|---|
| Residue: | 805 |
|---|
| Sequence: | MEPVMSLALAAHGPPSILEPLFKTVTTSTTTTTTTTTSTTTTTASPAGYSPGYPGTTLLT
ALFENLTSTAASGLYDPYSGMYGNQTNGTIGFETKGPRYSLASMVVMGFVAAILSTVTVA
GNVMVMISFKIDKQLQTISNYFLFSLAIADFAIGAISMPLFAVTTILGYWPLGPIVCDTW
LALDYLASNASVLNLLIISFDRYFSVTRPLTYRAKRTTNRAAVMIGAAWGISLLLWPPWI
YSWPYIEGKRTVPKDECYIQFIETNQYITFGTALAAFYFPVTIMCFLYWRIWRETKKRQK
DLPNLQAGKKDSSKRSNSSDENTVVNHASGGLLAFAQVGGNDHDTWRRPRSESSPDAESV
YMTNMVIDSGYHGMHSRKSSIKSTNTIKKSYTCFGSIKEWCIAWWHSGREDSDDFAYEQE
EPSDLGYATPVTIETPLQSSVSRCTSMNVMRDNYSMGGSVSGVRPPSILLSDVSPTPLPR
PPLASISQLQEMSAVTASTTANVNTSGNGNGAINNNNNASHNGNGAVNGNGAGNGSGIGL
GTTGNATHRDSRTLPVINRINSRSVSQDSVYTILIRLPSDGASSNAANGGGGGPGAGAAA
SASLSMQGDCAPSIKMIHEDGPTTTAAAAPLASAAATRRPLPSRDSEFSLPLGRRMSHAQ
HDARLLNAKVIPKQLGKAGGGAAGGGVGGAHALMNARNAAKKKKKSQEKRQESKAAKTLS
AILLSFIITWTPYNILVLIKPLTTCSDCIPTELWDFFYALCYINSTINPMCYALCNATFR
RTYVRILTCKWHTRNREGMVRGVYN
|
|
|
|---|
| BDBM50064176 |
|---|
| n/a |
|---|
| Name | BDBM50064176 |
|---|
| Synonyms: | CHEMBL27673 | CHEMBL500996 | METHOCTRAMINE | N,N''-Bis-[6-(2-methoxy-benzylamino)-hexyl]-octane-1,8-diamine | N1,N8-bis(6-(2-methoxybenzylamino)hexyl)octane-1,8-diamine |
|---|
| Type | Small organic molecule |
|---|
| Emp. Form. | C36H62N4O2 |
|---|
| Mol. Mass. | 582.9031 |
|---|
| SMILES | COc1ccccc1CNCCCCCCNCCCCCCCCNCCCCCCNCc1ccccc1OC |
|---|
| Structure | 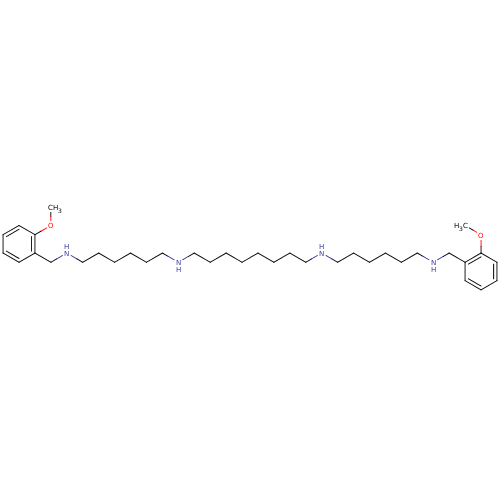
|
|---|


 Search and Browse
Search and Browse
 Download
Download
 Enter Data
Enter Data

 Honda, H; Tomizawa, M; Casida, JE Insect muscarinic acetylcholine receptor: pharmacological and toxicological profiles of antagonists and agonists. J Agric Food Chem55:2276-81 (2007) [PubMed] Article
Honda, H; Tomizawa, M; Casida, JE Insect muscarinic acetylcholine receptor: pharmacological and toxicological profiles of antagonists and agonists. J Agric Food Chem55:2276-81 (2007) [PubMed] Article