

 Search and Browse
Search and Browse
 Download
Download
 Enter Data
Enter Data
| Reaction Details | |||
|---|---|---|---|
 | Report a problem with these data | ||
| Target | Corticotropin-releasing factor receptor 1 | ||
| Ligand | BDBM50058163 | ||
| Substrate/Competitor | n/a | ||
| Meas. Tech. | ChEMBL_51114 (CHEMBL664960) | ||
| Ki | 1.61±n/a nM | ||
| Citation |  Chorvat, RJ; Bakthavatchalam, R; Beck, JP; Gilligan, PJ; Wilde, RG; Cocuzza, AJ; Hobbs, FW; Cheeseman, RS; Curry, M; Rescinito, JP; Krenitsky, P; Chidester, D; Yarem, JA; Klaczkiewicz, JD; Hodge, CN; Aldrich, PE; Wasserman, ZR; Fernandez, CH; Zaczek, R; Fitzgerald, LW; Huang, SM; Shen, HL; Wong, YN; Chien, BM; Arvanitis, A Synthesis, corticotropin-releasing factor receptor binding affinity, and pharmacokinetic properties of triazolo-, imidazo-, and pyrrolopyrimidines and -pyridines. J Med Chem42:833-48 (1999) [PubMed] Article Chorvat, RJ; Bakthavatchalam, R; Beck, JP; Gilligan, PJ; Wilde, RG; Cocuzza, AJ; Hobbs, FW; Cheeseman, RS; Curry, M; Rescinito, JP; Krenitsky, P; Chidester, D; Yarem, JA; Klaczkiewicz, JD; Hodge, CN; Aldrich, PE; Wasserman, ZR; Fernandez, CH; Zaczek, R; Fitzgerald, LW; Huang, SM; Shen, HL; Wong, YN; Chien, BM; Arvanitis, A Synthesis, corticotropin-releasing factor receptor binding affinity, and pharmacokinetic properties of triazolo-, imidazo-, and pyrrolopyrimidines and -pyridines. J Med Chem42:833-48 (1999) [PubMed] Article | ||
| More Info.: | Get all data from this article, Assay Method | ||
| Corticotropin-releasing factor receptor 1 | |||
| Name: | Corticotropin-releasing factor receptor 1 | ||
| Synonyms: | CRF-R | CRF-R2 Alpha | CRF1 | CRFR | CRFR1 | CRFR1_HUMAN | CRH-R 1 | CRHR | CRHR1 | Corticotropin releasing factor receptor 1 | Corticotropin-releasing factor receptor 1 (CRF-1) | Corticotropin-releasing factor receptor 1 (CRF1) | Corticotropin-releasing hormone receptor 1 | ||
| Type: | Enzyme | ||
| Mol. Mass.: | 50744.31 | ||
| Organism: | Homo sapiens (Human) | ||
| Description: | P34998 | ||
| Residue: | 444 | ||
| Sequence: |
| ||
| BDBM50058163 | |||
| n/a | |||
| Name | BDBM50058163 | ||
| Synonyms: | Butyl-[2,5-dimethyl-7-(2,4,6-trimethyl-phenyl)-7H-pyrrolo[2,3-d]pyrimidin-4-yl]-ethyl-amine | CHEMBL9946 | CP 154,526 | CP-154526 | N-butyl-N-ethyl-7-mesityl-2,5-dimethyl-7H-pyrrolo[2,3-d]pyrimidin-4-amine | ||
| Type | Small organic molecule | ||
| Emp. Form. | C23H32N4 | ||
| Mol. Mass. | 364.527 | ||
| SMILES | CCCCN(CC)c1nc(C)nc2n(cc(C)c12)-c1c(C)cc(C)cc1C |(-10.62,2.69,;-9.29,3.46,;-7.95,2.7,;-6.62,3.48,;-5.28,2.71,;-3.95,3.49,;-3.96,5.03,;-5.27,1.17,;-6.6,.4,;-6.6,-1.14,;-7.94,-1.91,;-5.27,-1.91,;-3.92,-1.14,;-2.44,-1.61,;-1.54,-.35,;-2.46,.9,;-1.99,2.37,;-3.93,.41,;-1.96,-3.07,;-2.99,-4.22,;-4.49,-3.91,;-2.5,-5.68,;-.99,-5.99,;-.51,-7.46,;.03,-4.83,;-.46,-3.38,;.56,-2.22,)| | ||
| Structure | 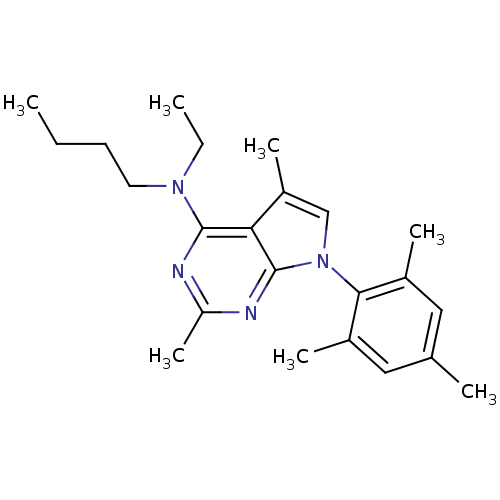
| ||