| Reaction Details |
|---|
 | Report a problem with these data |
| Target | Muscarinic acetylcholine receptor M3 |
|---|
| Ligand | BDBM50109647 |
|---|
| Substrate/Competitor | n/a |
|---|
| Meas. Tech. | ChEMBL_1972414 (CHEMBL4605232) |
|---|
| EC50 | 45±n/a nM |
|---|
| Citation |  Fischer, O; Hofmann, J; Rampp, H; Kaindl, J; Pratsch, G; Bartuschat, A; Taudte, RV; Fromm, MF; H�bner, H; Gmeiner, P; Heinrich, MR Regiospecific Introduction of Halogens on the 2-Aminobiphenyl Subunit Leading to Highly Potent and Selective M3 Muscarinic Acetylcholine Receptor Antagonists and Weak Inverse Agonists. J Med Chem63:4349-4369 (2020) [PubMed] Article Fischer, O; Hofmann, J; Rampp, H; Kaindl, J; Pratsch, G; Bartuschat, A; Taudte, RV; Fromm, MF; H�bner, H; Gmeiner, P; Heinrich, MR Regiospecific Introduction of Halogens on the 2-Aminobiphenyl Subunit Leading to Highly Potent and Selective M3 Muscarinic Acetylcholine Receptor Antagonists and Weak Inverse Agonists. J Med Chem63:4349-4369 (2020) [PubMed] Article |
|---|
| More Info.: | Get all data from this article, Assay Method |
|---|
| |
| Muscarinic acetylcholine receptor M3 |
|---|
| Name: | Muscarinic acetylcholine receptor M3 |
|---|
| Synonyms: | ACM3_HUMAN | CHRM3 | Cholinergic, muscarinic M3 | Muscarinic Receptors M3 | Muscarinic receptor M3 | RecName: Full=Muscarinic acetylcholine receptor M3 |
|---|
| Type: | Enzyme |
|---|
| Mol. Mass.: | 66151.03 |
|---|
| Organism: | Homo sapiens (Human) |
|---|
| Description: | P20309 |
|---|
| Residue: | 590 |
|---|
| Sequence: | MTLHNNSTTSPLFPNISSSWIHSPSDAGLPPGTVTHFGSYNVSRAAGNFSSPDGTTDDPL
GGHTVWQVVFIAFLTGILALVTIIGNILVIVSFKVNKQLKTVNNYFLLSLACADLIIGVI
SMNLFTTYIIMNRWALGNLACDLWLAIDYVASNASVMNLLVISFDRYFSITRPLTYRAKR
TTKRAGVMIGLAWVISFVLWAPAILFWQYFVGKRTVPPGECFIQFLSEPTITFGTAIAAF
YMPVTIMTILYWRIYKETEKRTKELAGLQASGTEAETENFVHPTGSSRSCSSYELQQQSM
KRSNRRKYGRCHFWFTTKSWKPSSEQMDQDHSSSDSWNNNDAAASLENSASSDEEDIGSE
TRAIYSIVLKLPGHSTILNSTKLPSSDNLQVPEEELGMVDLERKADKLQAQKSVDDGGSF
PKSFSKLPIQLESAVDTAKTSDVNSSVGKSTATLPLSFKEATLAKRFALKTRSQITKRKR
MSLVKEKKAAQTLSAILLAFIITWTPYNIMVLVNTFCDSCIPKTFWNLGYWLCYINSTVN
PVCYALCNKTFRTTFKMLLLCQCDKKKRRKQQYQQRQSVIFHKRAPEQAL
|
|
|
|---|
| BDBM50109647 |
|---|
| n/a |
|---|
| Name | BDBM50109647 |
|---|
| Synonyms: | 2-{1-[2-(2,3-Dihydro-benzofuran-5-yl)-ethyl]-pyrrolidin-3-yl}-2,2-diphenyl-acetamide (Darifenacin) | CHEMBL1346 | DARIFENACIN | DARIFENACIN HYDROBROMIDE | Enablex |
|---|
| Type | Small organic molecule |
|---|
| Emp. Form. | C28H30N2O2 |
|---|
| Mol. Mass. | 426.55 |
|---|
| SMILES | NC(=O)C([C@@H]1CCN(CCc2ccc3OCCc3c2)C1)(c1ccccc1)c1ccccc1 |
|---|
| Structure | 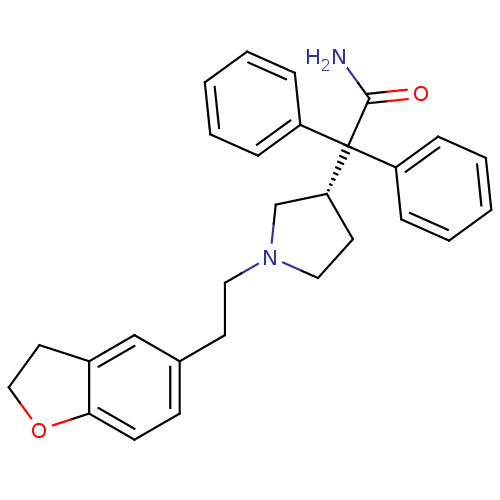
|
|---|


 Search and Browse
Search and Browse
 Download
Download
 Enter Data
Enter Data

 Fischer, O; Hofmann, J; Rampp, H; Kaindl, J; Pratsch, G; Bartuschat, A; Taudte, RV; Fromm, MF; H�bner, H; Gmeiner, P; Heinrich, MR Regiospecific Introduction of Halogens on the 2-Aminobiphenyl Subunit Leading to Highly Potent and Selective M3 Muscarinic Acetylcholine Receptor Antagonists and Weak Inverse Agonists. J Med Chem63:4349-4369 (2020) [PubMed] Article
Fischer, O; Hofmann, J; Rampp, H; Kaindl, J; Pratsch, G; Bartuschat, A; Taudte, RV; Fromm, MF; H�bner, H; Gmeiner, P; Heinrich, MR Regiospecific Introduction of Halogens on the 2-Aminobiphenyl Subunit Leading to Highly Potent and Selective M3 Muscarinic Acetylcholine Receptor Antagonists and Weak Inverse Agonists. J Med Chem63:4349-4369 (2020) [PubMed] Article