| Reaction Details |
|---|
 | Report a problem with these data |
| Target | Protein kinase C delta type |
|---|
| Ligand | BDBM50057514 |
|---|
| Substrate/Competitor | n/a |
|---|
| Meas. Tech. | ChEMBL_161472 (CHEMBL770275) |
|---|
| Ki | 0.1±n/a nM |
|---|
| Citation |  Rong, SB; Enyedy, IJ; Qiao, L; Zhao, L; Ma, D; Pearce, LL; Lorenzo, PS; Stone, JC; Blumberg, PM; Wang, S; Kozikowski, AP Structural basis of RasGRP binding to high-affinity PKC ligands. J Med Chem45:853-60 (2002) [PubMed] Rong, SB; Enyedy, IJ; Qiao, L; Zhao, L; Ma, D; Pearce, LL; Lorenzo, PS; Stone, JC; Blumberg, PM; Wang, S; Kozikowski, AP Structural basis of RasGRP binding to high-affinity PKC ligands. J Med Chem45:853-60 (2002) [PubMed] |
|---|
| More Info.: | Get all data from this article, Assay Method |
|---|
| |
| Protein kinase C delta type |
|---|
| Name: | Protein kinase C delta type |
|---|
| Synonyms: | KPCD_MOUSE | Pkcd | Prkcd | Protein kinase C delta | Protein kinase C delta type | nPKC-delta |
|---|
| Type: | PROTEIN |
|---|
| Mol. Mass.: | 77554.99 |
|---|
| Organism: | Mus musculus |
|---|
| Description: | ChEMBL_1351519 |
|---|
| Residue: | 674 |
|---|
| Sequence: | MAPFLRISFNSYELGSLQVEDEASQPFCAVKMKEALSTERGKTLVQKKPTMYPEWKTTFD
AHIYEGRVIQIVLMRAAEDPVSEVTVGVSVLAERCKKNNGKAEFWLDLQPQAKVLMCVQY
FLEDGDCKQSMRSEEEAKFPTMNRRGAIKQAKIHYIKNHEFIATFFGQPTFCSVCKEFVW
GLNKQGYKCRQCNAAIHKKCIDKIIGRCTGTATNSRDTIFQKERFNIDMPHRFKVYNYMS
PTFCDHCGSLLWGLVKQGLKCEDCGMNVHHKCREKVANLCGINQKLLAEALNQVTQRSSR
KLDTTESVGIYQGFEKKPEVSGSDILDNNGTYGKIWEGSTRCTLENFTFQKVLGKGSFGK
VLLAELKGKDKYFAIKCLKKDVVLIDDDVECTMVEKRVLALAWESPFLTHLICTFQTKDH
LFFVMEFLNGGDLMFHIQDKGRFELYRATFYAAEIICGLQFLHSKGIIYRDLKLDNVMLD
RDGHIKIADFGMCKENIFGEGRASTFCGTPDYIAPEILQGLKYSFSVDWWSFGVLLYEML
IGQSPFHGDDEDELFESIRVDTPHYPRWITKESKDIMEKLFERDPDKRLGVTGNIRIHPF
FKTINWSLLEKRKVEPPFKPKVKSPSDYSNFDPEFLNEKPQLSFSDKNLIDSMDQEAFHG
FSFVNPKFEQFLDI
|
|
|
|---|
| BDBM50057514 |
|---|
| n/a |
|---|
| Name | BDBM50057514 |
|---|
| Synonyms: | (10S,13S)-13-Hydroxymethyl-10-isopropyl-9-methyl-5-octyl-3,9,12-triaza-tricyclo[6.6.1.0*4,15*]pentadeca-1,4,6,8(15)-tetraen-11-one | 13-Hydroxymethyl-10-isopropyl-9-methyl-5-octyl-3,9,12-triaza-tricyclo[6.6.1.0*4,15*]pentadeca-1,4,6,8(15)-tetraen-11-one (n-octyl-ILV) | CHEMBL285801 |
|---|
| Type | Small organic molecule |
|---|
| Emp. Form. | C25H39N3O2 |
|---|
| Mol. Mass. | 413.5961 |
|---|
| SMILES | CCCCCCCCc1ccc2N(C)[C@@H](C(C)C)C(=O)N[C@H](CO)Cc3c[nH]c1c23 |
|---|
| Structure | 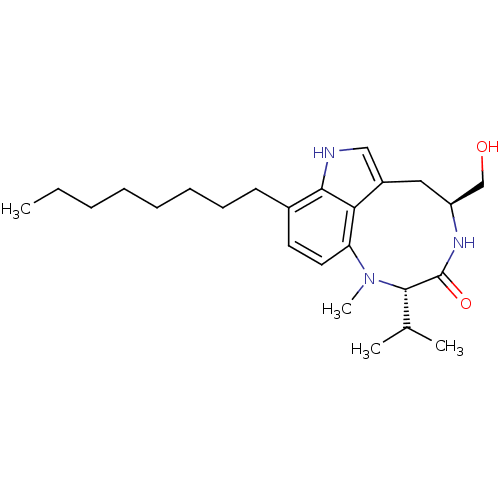
|
|---|


 Search and Browse
Search and Browse
 Download
Download
 Enter Data
Enter Data

 Rong, SB; Enyedy, IJ; Qiao, L; Zhao, L; Ma, D; Pearce, LL; Lorenzo, PS; Stone, JC; Blumberg, PM; Wang, S; Kozikowski, AP Structural basis of RasGRP binding to high-affinity PKC ligands. J Med Chem45:853-60 (2002) [PubMed]
Rong, SB; Enyedy, IJ; Qiao, L; Zhao, L; Ma, D; Pearce, LL; Lorenzo, PS; Stone, JC; Blumberg, PM; Wang, S; Kozikowski, AP Structural basis of RasGRP binding to high-affinity PKC ligands. J Med Chem45:853-60 (2002) [PubMed]