| Reaction Details |
|---|
 | Report a problem with these data |
| Target | Poly [ADP-ribose] polymerase 1 |
|---|
| Ligand | BDBM50165488 |
|---|
| Substrate/Competitor | n/a |
|---|
| Meas. Tech. | ChEMBL_305621 (CHEMBL829484) |
|---|
| IC50 | 4.1±n/a nM |
|---|
| Citation |  Loh, VM; Cockcroft, XL; Dillon, KJ; Dixon, L; Drzewiecki, J; Eversley, PJ; Gomez, S; Hoare, J; Kerrigan, F; Matthews, IT; Menear, KA; Martin, NM; Newton, RF; Paul, J; Smith, GC; Vile, J; Whittle, AJ Phthalazinones. Part 1: The design and synthesis of a novel series of potent inhibitors of poly(ADP-ribose)polymerase. Bioorg Med Chem Lett15:2235-8 (2005) [PubMed] Article Loh, VM; Cockcroft, XL; Dillon, KJ; Dixon, L; Drzewiecki, J; Eversley, PJ; Gomez, S; Hoare, J; Kerrigan, F; Matthews, IT; Menear, KA; Martin, NM; Newton, RF; Paul, J; Smith, GC; Vile, J; Whittle, AJ Phthalazinones. Part 1: The design and synthesis of a novel series of potent inhibitors of poly(ADP-ribose)polymerase. Bioorg Med Chem Lett15:2235-8 (2005) [PubMed] Article |
|---|
| More Info.: | Get all data from this article, Assay Method |
|---|
| |
| Poly [ADP-ribose] polymerase 1 |
|---|
| Name: | Poly [ADP-ribose] polymerase 1 |
|---|
| Synonyms: | (ARTD1 or PARP1) | 2.4.2.- | 2.4.2.30 | ADP-ribosyltransferase diphtheria toxin-like 1 | ADPRT | ADPRT 1 | ARTD1 | DNA ADP-ribosyltransferase PARP1 | Human diphtheria toxin-like ADP-ribosyltransferase (ARTD1 or PARP1) | NAD(+) ADP-ribosyltransferase 1 | NT-PARP-1 | PARP-1 | PARP1 | PARP1_HUMAN | PPOL | Poly [ADP-ribose] polymerase (PARP) | Poly [ADP-ribose] polymerase 1 (PARP) | Poly [ADP-ribose] polymerase 1 (PARP-1) | Poly [ADP-ribose] polymerase 1 (PARP1) | Poly [ADP-ribose] polymerase 1, 24-kDa form | Poly [ADP-ribose] polymerase 1, 28-kDa form | Poly [ADP-ribose] polymerase 1, 89-kDa form | Poly [ADP-ribose] polymerase 1, processed C-terminus | Poly [ADP-ribose] polymerase 1, processed N-terminus | Poly [ADP-ribose] polymerase-1 | Poly(ADP-ribose) polymerase 1 (PARP1) | Poly(ADP-ribose) polymerase-1 (ARTD1/PARP1) | Poly[ADP-ribose] synthase 1 | Protein poly-ADP-ribosyltransferase PARP1 | Synonyms=ADPRT |
|---|
| Type: | n/a |
|---|
| Mol. Mass.: | 113114.22 |
|---|
| Organism: | Homo sapiens (Human) |
|---|
| Description: | P09874 |
|---|
| Residue: | 1014 |
|---|
| Sequence: | MAESSDKLYRVEYAKSGRASCKKCSESIPKDSLRMAIMVQSPMFDGKVPHWYHFSCFWKV
GHSIRHPDVEVDGFSELRWDDQQKVKKTAEAGGVTGKGQDGIGSKAEKTLGDFAAEYAKS
NRSTCKGCMEKIEKGQVRLSKKMVDPEKPQLGMIDRWYHPGCFVKNREELGFRPEYSASQ
LKGFSLLATEDKEALKKQLPGVKSEGKRKGDEVDGVDEVAKKKSKKEKDKDSKLEKALKA
QNDLIWNIKDELKKVCSTNDLKELLIFNKQQVPSGESAILDRVADGMVFGALLPCEECSG
QLVFKSDAYYCTGDVTAWTKCMVKTQTPNRKEWVTPKEFREISYLKKLKVKKQDRIFPPE
TSASVAATPPPSTASAPAAVNSSASADKPLSNMKILTLGKLSRNKDEVKAMIEKLGGKLT
GTANKASLCISTKKEVEKMNKKMEEVKEANIRVVSEDFLQDVSASTKSLQELFLAHILSP
WGAEVKAEPVEVVAPRGKSGAALSKKSKGQVKEEGINKSEKRMKLTLKGGAAVDPDSGLE
HSAHVLEKGGKVFSATLGLVDIVKGTNSYYKLQLLEDDKENRYWIFRSWGRVGTVIGSNK
LEQMPSKEDAIEHFMKLYEEKTGNAWHSKNFTKYPKKFYPLEIDYGQDEEAVKKLTVNPG
TKSKLPKPVQDLIKMIFDVESMKKAMVEYEIDLQKMPLGKLSKRQIQAAYSILSEVQQAV
SQGSSDSQILDLSNRFYTLIPHDFGMKKPPLLNNADSVQAKVEMLDNLLDIEVAYSLLRG
GSDDSSKDPIDVNYEKLKTDIKVVDRDSEEAEIIRKYVKNTHATTHNAYDLEVIDIFKIE
REGECQRYKPFKQLHNRRLLWHGSRTTNFAGILSQGLRIAPPEAPVTGYMFGKGIYFADM
VSKSANYCHTSQGDPIGLILLGEVALGNMYELKHASHISKLPKGKHSVKGLGKTTPDPSA
NISLDGVDVPLGTGISSGVNDTSLLYNEYIVYDIAQVNLKYLLKLKFNFKTSLW
|
|
|
|---|
| BDBM50165488 |
|---|
| n/a |
|---|
| Name | BDBM50165488 |
|---|
| Synonyms: | 1-(2-fluoro-5-((4-oxo-3,4-dihydrophthalazin-1-yl)methyl)phenyl)piperidine-2,6-dione | 1-[2-Fluoro-5-(4-oxo-3,4-dihydro-phthalazin-1-ylmethyl)-phenyl]-piperidine-2,6-dione | CHEMBL370217 |
|---|
| Type | Small organic molecule |
|---|
| Emp. Form. | C20H16FN3O3 |
|---|
| Mol. Mass. | 365.3577 |
|---|
| SMILES | Fc1ccc(Cc2n[nH]c(=O)c3ccccc23)cc1N1C(=O)CCCC1=O |
|---|
| Structure | 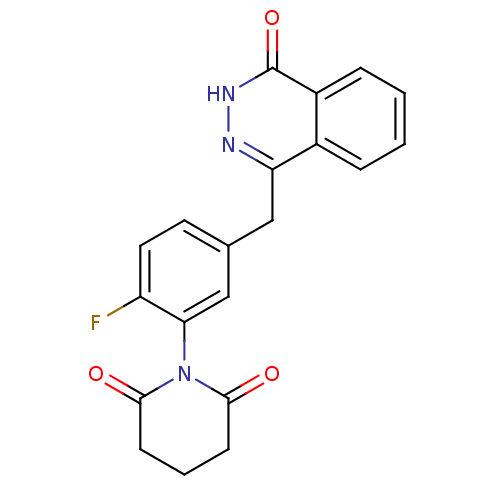
|
|---|


 Search and Browse
Search and Browse
 Download
Download
 Enter Data
Enter Data

 Loh, VM; Cockcroft, XL; Dillon, KJ; Dixon, L; Drzewiecki, J; Eversley, PJ; Gomez, S; Hoare, J; Kerrigan, F; Matthews, IT; Menear, KA; Martin, NM; Newton, RF; Paul, J; Smith, GC; Vile, J; Whittle, AJ Phthalazinones. Part 1: The design and synthesis of a novel series of potent inhibitors of poly(ADP-ribose)polymerase. Bioorg Med Chem Lett15:2235-8 (2005) [PubMed] Article
Loh, VM; Cockcroft, XL; Dillon, KJ; Dixon, L; Drzewiecki, J; Eversley, PJ; Gomez, S; Hoare, J; Kerrigan, F; Matthews, IT; Menear, KA; Martin, NM; Newton, RF; Paul, J; Smith, GC; Vile, J; Whittle, AJ Phthalazinones. Part 1: The design and synthesis of a novel series of potent inhibitors of poly(ADP-ribose)polymerase. Bioorg Med Chem Lett15:2235-8 (2005) [PubMed] Article