| Reaction Details |
|---|
 | Report a problem with these data |
| Target | Cannabinoid receptor 1 |
|---|
| Ligand | BDBM50176988 |
|---|
| Substrate/Competitor | n/a |
|---|
| Meas. Tech. | ChEMBL_430951 (CHEMBL914893) |
|---|
| Ki | 0.00035±n/a nM |
|---|
| Citation |  Murineddu, G; Ruiu, S; Loriga, G; Manca, I; Lazzari, P; Reali, R; Pani, L; Toma, L; Pinna, GA Tricyclic pyrazoles. 3. Synthesis, biological evaluation, and molecular modeling of analogues of the cannabinoid antagonist 8-chloro-1-(2',4'-dichlorophenyl)-N-piperidin-1-yl-1,4,5,6-tetrahydrobenzo[6,7]cyclohepta[1,2-c]pyrazole-3-carboxamide. J Med Chem48:7351-62 (2005) [PubMed] Article Murineddu, G; Ruiu, S; Loriga, G; Manca, I; Lazzari, P; Reali, R; Pani, L; Toma, L; Pinna, GA Tricyclic pyrazoles. 3. Synthesis, biological evaluation, and molecular modeling of analogues of the cannabinoid antagonist 8-chloro-1-(2',4'-dichlorophenyl)-N-piperidin-1-yl-1,4,5,6-tetrahydrobenzo[6,7]cyclohepta[1,2-c]pyrazole-3-carboxamide. J Med Chem48:7351-62 (2005) [PubMed] Article |
|---|
| More Info.: | Get all data from this article, Assay Method |
|---|
| |
| Cannabinoid receptor 1 |
|---|
| Name: | Cannabinoid receptor 1 |
|---|
| Synonyms: | Brain-type cannabinoid receptor | CANNABINOID CB1 | CB-R | CB1 | CNR1_MOUSE | Cannabinoid CB1 receptor | Cannabinoid receptor | Cnr1 |
|---|
| Type: | Enzyme |
|---|
| Mol. Mass.: | 52842.52 |
|---|
| Organism: | Mus musculus (Mouse) |
|---|
| Description: | P47746 |
|---|
| Residue: | 473 |
|---|
| Sequence: | MKSILDGLADTTFRTITTDLLYVGSNDIQYEDIKGDMASKLGYFPQKFPLTSFRGSPFQE
KMTAGDNSPLVPAGDTTNITEFYNKSLSSFKENEDNIQCGENFMDMECFMILNPSQQLAI
AVLSLTLGTFTVLENLLVLCVILHSRSLRCRPSYHFIGSLAVADLLGSVIFVYSFVDFHV
FHRKDSPNVFLFKLGGVTASFTASVGSLFLTAIDRYISIHRPLAYKRIVTRPKAVVAFCL
MWTIAIVIAVLPLLGWNCKKLQSVCSDIFPLIDETYLMFWIGVTSVLLLFIVYAYMYILW
KAHSHAVRMIQRGTQKSIIIHTSEDGKVQVTRPDQARMDIRLAKTLVLILVVLIICWGPL
LAIMVYDVFGKMNKLIKTVFAFCSMLCLLNSTVNPIIYALRSKDLRHAFRSMFPSCEGTA
QPLDNSMGDSDCLHKHANNTASMHRAAESCIKSTVKIAKVTMSVSTDTSAEAL
|
|
|
|---|
| BDBM50176988 |
|---|
| n/a |
|---|
| Name | BDBM50176988 |
|---|
| Synonyms: | 8-Chloro-1-(2,4-dichloro-phenyl)-1,3a,4,5,6,10b-hexahydro-1,2-diaza-benzo[e]azulene-3-carboxylic acid piperidin-1-ylamide | 8-chloro-1-(2',4'-dichlorophenyl)-N-piperidin-1-yl-1,4,5,6-tetrahydrobenzo[6,7]cyclohepta[1,2-c]pyrazole-3-carboxamide | 8-chloro-1-(2',4'-dichlorophenyl)-N-piperidin-1-yl-1,4,5,6-tetrahydrobenzo[6,7]cyclohepta[1,2-c]pyrazole-3-carboxamide | 8-chloro-1-(2,4-dichloro-phenyl)-1,4,5,6-tetrahydro-1,2-diaza-benzo[e]azulene-3-carboxylic acid piperidin-1-ylamide | CHEMBL376700 |
|---|
| Type | Small organic molecule |
|---|
| Emp. Form. | C24H23Cl3N4O |
|---|
| Mol. Mass. | 489.825 |
|---|
| SMILES | Clc1ccc(c(Cl)c1)-n1nc(C(=O)NN2CCCCC2)c2CCCc3cc(Cl)ccc3-c12 |
|---|
| Structure | 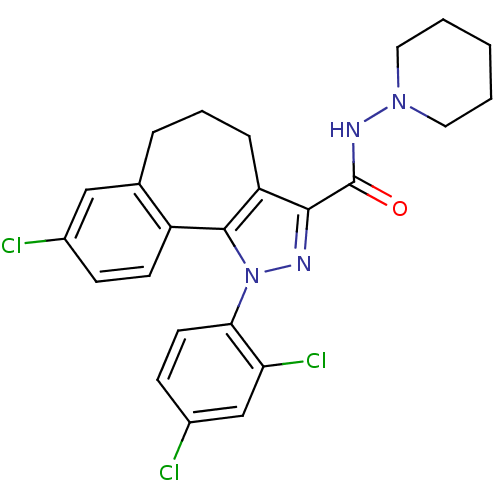
|
|---|


 Search and Browse
Search and Browse
 Download
Download
 Enter Data
Enter Data

 Murineddu, G; Ruiu, S; Loriga, G; Manca, I; Lazzari, P; Reali, R; Pani, L; Toma, L; Pinna, GA Tricyclic pyrazoles. 3. Synthesis, biological evaluation, and molecular modeling of analogues of the cannabinoid antagonist 8-chloro-1-(2',4'-dichlorophenyl)-N-piperidin-1-yl-1,4,5,6-tetrahydrobenzo[6,7]cyclohepta[1,2-c]pyrazole-3-carboxamide. J Med Chem48:7351-62 (2005) [PubMed] Article
Murineddu, G; Ruiu, S; Loriga, G; Manca, I; Lazzari, P; Reali, R; Pani, L; Toma, L; Pinna, GA Tricyclic pyrazoles. 3. Synthesis, biological evaluation, and molecular modeling of analogues of the cannabinoid antagonist 8-chloro-1-(2',4'-dichlorophenyl)-N-piperidin-1-yl-1,4,5,6-tetrahydrobenzo[6,7]cyclohepta[1,2-c]pyrazole-3-carboxamide. J Med Chem48:7351-62 (2005) [PubMed] Article