| Reaction Details |
|---|
 | Report a problem with these data |
| Target | Delta-type opioid receptor |
|---|
| Ligand | BDBM50252417 |
|---|
| Substrate/Competitor | n/a |
|---|
| Meas. Tech. | ChEMBL_487721 (CHEMBL1009453) |
|---|
| Ki | 4228±n/a nM |
|---|
| Citation |  Keresztes, A; Szucs, M; Borics, A; Kövér, KE; Forró, E; Fülöp, F; Tömböly, C; Péter, A; Páhi, A; Fábián, G; Murányi, M; Tóth, G New endomorphin analogues containing alicyclic beta-amino acids: influence on bioactive conformation and pharmacological profile. J Med Chem51:4270-9 (2008) [PubMed] Article Keresztes, A; Szucs, M; Borics, A; Kövér, KE; Forró, E; Fülöp, F; Tömböly, C; Péter, A; Páhi, A; Fábián, G; Murányi, M; Tóth, G New endomorphin analogues containing alicyclic beta-amino acids: influence on bioactive conformation and pharmacological profile. J Med Chem51:4270-9 (2008) [PubMed] Article |
|---|
| More Info.: | Get all data from this article, Assay Method |
|---|
| |
| Delta-type opioid receptor |
|---|
| Name: | Delta-type opioid receptor |
|---|
| Synonyms: | Cytochrome P450 3A4 | DOR-1 | Delta opioid receptor | Delta-type opioid receptor | Delta-type opioid receptor (DOR) | OPIATE Delta | OPRD_RAT | Opiate Delta 1 | Opioid receptor | Opioid receptor A | Opioid receptors; mu & delta | Oprd1 | Ror-a | Voltage-gated potassium channel |
|---|
| Type: | G Protein-Coupled Receptor (GPCR) |
|---|
| Mol. Mass.: | 40465.04 |
|---|
| Organism: | Rattus norvegicus (rat) |
|---|
| Description: | Competition binding assays were using CHO-K1 cell membranes expressing the opioid receptor. |
|---|
| Residue: | 372 |
|---|
| Sequence: | MEPVPSARAELQFSLLANVSDTFPSAFPSASANASGSPGARSASSLALAIAITALYSAVC
AVGLLGNVLVMFGIVRYTKLKTATNIYIFNLALADALATSTLPFQSAKYLMETWPFGELL
CKAVLSIDYYNMFTSIFTLTMMSVDRYIAVCHPVKALDFRTPAKAKLINICIWVLASGVG
VPIMVMAVTQPRDGAVVCTLQFPSPSWYWDTVTKICVFLFAFVVPILIITVCYGLMLLRL
RSVRLLSGSKEKDRSLRRITRMVLVVVGAFVVCWAPIHIFVIVWTLVDINRRDPLVVAAL
HLCIALGYANSSLNPVLYAFLDENFKRCFRQLCRAPCGGQEPGSLRRPRQATARERVTAC
TPSDGPGGGAAA
|
|
|
|---|
| BDBM50252417 |
|---|
| n/a |
|---|
| Name | BDBM50252417 |
|---|
| Synonyms: | (1S,2S)-N-((S)-1-((S)-1-amino-1-oxo-3-phenylpropan-2-ylamino)-1-oxo-3-phenylpropan-2-yl)-2-((S)-2-amino-3-(4-hydroxyphenyl)propanamido)cyclopentanecarboxamide | CHEMBL467089 |
|---|
| Type | Small organic molecule |
|---|
| Emp. Form. | C33H39N5O5 |
|---|
| Mol. Mass. | 585.6933 |
|---|
| SMILES | N[C@@H](Cc1ccc(O)cc1)C(=O)N[C@H]1CCC[C@@H]1C(=O)N[C@@H](Cc1ccccc1)C(=O)N[C@@H](Cc1ccccc1)C(N)=O |r| |
|---|
| Structure | 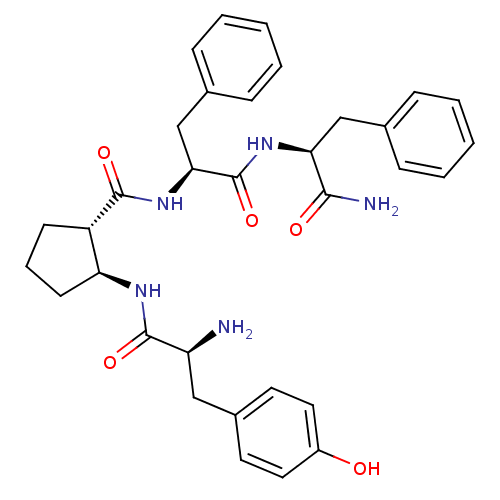
|
|---|


 Search and Browse
Search and Browse
 Download
Download
 Enter Data
Enter Data

 Keresztes, A; Szucs, M; Borics, A; Kövér, KE; Forró, E; Fülöp, F; Tömböly, C; Péter, A; Páhi, A; Fábián, G; Murányi, M; Tóth, G New endomorphin analogues containing alicyclic beta-amino acids: influence on bioactive conformation and pharmacological profile. J Med Chem51:4270-9 (2008) [PubMed] Article
Keresztes, A; Szucs, M; Borics, A; Kövér, KE; Forró, E; Fülöp, F; Tömböly, C; Péter, A; Páhi, A; Fábián, G; Murányi, M; Tóth, G New endomorphin analogues containing alicyclic beta-amino acids: influence on bioactive conformation and pharmacological profile. J Med Chem51:4270-9 (2008) [PubMed] Article