

 Search and Browse
Search and Browse
 Download
Download
 Enter Data
Enter Data
| Reaction Details | |||
|---|---|---|---|
 | Report a problem with these data | ||
| Target | Integrin alpha-L | ||
| Ligand | BDBM50265285 | ||
| Substrate/Competitor | n/a | ||
| Meas. Tech. | ChEMBL_497066 (CHEMBL997702) | ||
| IC50 | 4±n/a nM | ||
| Citation |  Lin, EY; Guckian, KM; Silvian, L; Chin, D; Boriack-Sjodin, PA; van Vlijmen, H; Friedman, JE; Scott, DM Structure-activity relationship of ortho- and meta-phenol based LFA-1 ICAM inhibitors. Bioorg Med Chem Lett18:5245-8 (2008) [PubMed] Article Lin, EY; Guckian, KM; Silvian, L; Chin, D; Boriack-Sjodin, PA; van Vlijmen, H; Friedman, JE; Scott, DM Structure-activity relationship of ortho- and meta-phenol based LFA-1 ICAM inhibitors. Bioorg Med Chem Lett18:5245-8 (2008) [PubMed] Article | ||
| More Info.: | Get all data from this article, Assay Method | ||
| Integrin alpha-L | |||
| Name: | Integrin alpha-L | ||
| Synonyms: | CD11 antigen-like family member A | CD11A | CD_antigen=CD11a | ITAL_HUMAN | ITGAL | Integrin alpha-L/beta-2 (LFA-1) | LFA-1A | Leukocyte adhesion glycoprotein LFA-1 alpha | Leukocyte adhesion glycoprotein LFA-1 alpha chain | Leukocyte function-associated molecule 1 alpha chain | ||
| Type: | PROTEIN | ||
| Mol. Mass.: | 128748.90 | ||
| Organism: | Homo sapiens (Human) | ||
| Description: | ChEMBL_629045 | ||
| Residue: | 1170 | ||
| Sequence: |
| ||
| BDBM50265285 | |||
| n/a | |||
| Name | BDBM50265285 | ||
| Synonyms: | (cis)-4-(2-(4-((E)-3-morpholino-3-oxoprop-1-enyl)-2,3-bis(trifluoromethyl)phenylthio)phenoxy)cyclohexanecarboxylic acid | CHEMBL502512 | ||
| Type | Small organic molecule | ||
| Emp. Form. | C28H27F6NO5S | ||
| Mol. Mass. | 603.573 | ||
| SMILES | OC(=O)[C@@H]1CC[C@@H](CC1)Oc1ccccc1Sc1ccc(\C=C\C(=O)N2CCOCC2)c(c1C(F)(F)F)C(F)(F)F |r,wU:6.9,3.2,(-5.15,-20.16,;-5.15,-21.7,;-6.48,-22.47,;-3.81,-22.48,;-3.81,-24.02,;-2.47,-24.79,;-1.13,-24.02,;-1.14,-22.47,;-2.47,-21.71,;.2,-24.79,;.21,-26.33,;-1.12,-27.1,;-1.12,-28.64,;.21,-29.41,;1.55,-28.64,;1.54,-27.09,;2.87,-26.32,;4.21,-27.08,;4.21,-28.62,;5.54,-29.38,;6.88,-28.61,;8.21,-29.37,;9.54,-28.59,;10.88,-29.36,;12.21,-28.58,;10.89,-30.9,;9.55,-31.66,;9.56,-33.19,;10.89,-33.97,;12.23,-33.19,;12.22,-31.65,;6.87,-27.06,;5.53,-26.3,;5.52,-24.76,;5.51,-23.22,;3.98,-24.77,;7.06,-24.75,;8.19,-26.28,;9.52,-25.5,;8.97,-27.61,;8.18,-24.73,)| | ||
| Structure | 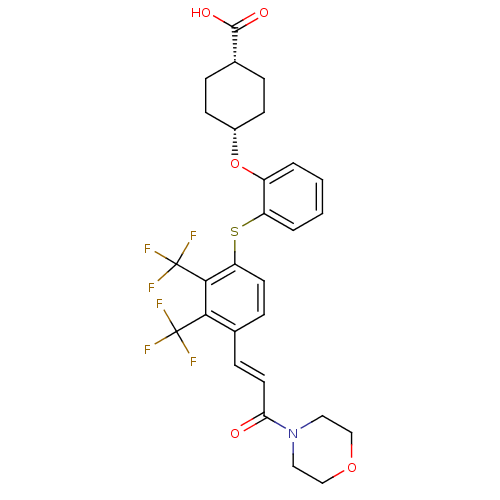
| ||