| Reaction Details |
|---|
 | Report a problem with these data |
| Target | Fatty-acid amide hydrolase 1 [30-579] |
|---|
| Ligand | BDBM50298650 |
|---|
| Substrate/Competitor | n/a |
|---|
| Meas. Tech. | ChEMBL_587912 (CHEMBL1046585) |
|---|
| IC50 | 3000±n/a nM |
|---|
| Citation |  Myllymäki, MJ; Käsnänen, H; Kataja, AO; Lahtela-Kakkonen, M; Saario, SM; Poso, A; Koskinen, AM Chiral 3-(4,5-dihydrooxazol-2-yl)phenyl alkylcarbamates as novel FAAH inhibitors: Insight into FAAH enantioselectivity by molecular docking and interaction fields. Eur J Med Chem44:4179-91 (2009) [PubMed] Article Myllymäki, MJ; Käsnänen, H; Kataja, AO; Lahtela-Kakkonen, M; Saario, SM; Poso, A; Koskinen, AM Chiral 3-(4,5-dihydrooxazol-2-yl)phenyl alkylcarbamates as novel FAAH inhibitors: Insight into FAAH enantioselectivity by molecular docking and interaction fields. Eur J Med Chem44:4179-91 (2009) [PubMed] Article |
|---|
| More Info.: | Get all data from this article, Assay Method |
|---|
| |
| Fatty-acid amide hydrolase 1 [30-579] |
|---|
| Name: | Fatty-acid amide hydrolase 1 [30-579] |
|---|
| Synonyms: | Anandamide amidohydrolase 1 | FAAH1_RAT | Faah | Faah1 | Fatty Acid Amide Hydrolase | Fatty Acid Amide Hydrolic, FAAH | Fatty-acid amide hydrolase (FAAH) | Fatty-acid amide hydrolase 1 | Fatty-acid amide hydrolase 1 (FAAH) | Fatty-acid amide hydrolase 1 (aa 30-579) | Oleamide hydrolase 1 |
|---|
| Type: | Single-pass membrane protein; homodimer |
|---|
| Mol. Mass.: | 60474.00 |
|---|
| Organism: | Rattus norvegicus (rat) |
|---|
| Description: | P97612 (aa 30-579) |
|---|
| Residue: | 550 |
|---|
| Sequence: | RWTGRQKARGAATRARQKQRASLETMDKAVQRFRLQNPDLDSEALLTLPLLQLVQKLQSG
ELSPEAVFFTYLGKAWEVNKGTNCVTSYLTDCETQLSQAPRQGLLYGVPVSLKECFSYKG
HDSTLGLSLNEGMPSESDCVVVQVLKLQGAVPFVHTNVPQSMLSFDCSNPLFGQTMNPWK
SSKSPGGSSGGEGALIGSGGSPLGLGTDIGGSIRFPSAFCGICGLKPTGNRLSKSGLKGC
VYGQTAVQLSLGPMARDVESLALCLKALLCEHLFTLDPTVPPLPFREEVYRSSRPLRVGY
YETDNYTMPSPAMRRALIETKQRLEAAGHTLIPFLPNNIPYALEVLSAGGLFSDGGRSFL
QNFKGDFVDPCLGDLILILRLPSWFKRLLSLLLKPLFPRLAAFLNSMRPRSAEKLWKLQH
EIEMYRQSVIAQWKAMNLDVLLTPMLGPALDLNTPGRATGAISYTVLYNCLDFPAGVVPV
TTVTAEDDAQMELYKGYFGDIWDIILKKAMKNSVGLPVAVQCVALPWQEELCLRFMREVE
QLMTPQKQPS
|
|
|
|---|
| BDBM50298650 |
|---|
| n/a |
|---|
| Name | BDBM50298650 |
|---|
| Synonyms: | 3-(benzo[d]oxazol-2-yl)phenyl propylcarbamate | CHEMBL573871 |
|---|
| Type | Small organic molecule |
|---|
| Emp. Form. | C17H16N2O3 |
|---|
| Mol. Mass. | 296.3205 |
|---|
| SMILES | CCCNC(=O)Oc1cccc(c1)-c1nc2ccccc2o1 |
|---|
| Structure | 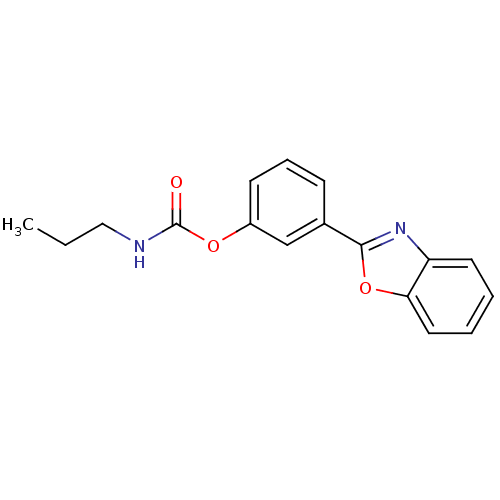
|
|---|


 Search and Browse
Search and Browse
 Download
Download
 Enter Data
Enter Data

 Myllymäki, MJ; Käsnänen, H; Kataja, AO; Lahtela-Kakkonen, M; Saario, SM; Poso, A; Koskinen, AM Chiral 3-(4,5-dihydrooxazol-2-yl)phenyl alkylcarbamates as novel FAAH inhibitors: Insight into FAAH enantioselectivity by molecular docking and interaction fields. Eur J Med Chem44:4179-91 (2009) [PubMed] Article
Myllymäki, MJ; Käsnänen, H; Kataja, AO; Lahtela-Kakkonen, M; Saario, SM; Poso, A; Koskinen, AM Chiral 3-(4,5-dihydrooxazol-2-yl)phenyl alkylcarbamates as novel FAAH inhibitors: Insight into FAAH enantioselectivity by molecular docking and interaction fields. Eur J Med Chem44:4179-91 (2009) [PubMed] Article