

 Search and Browse
Search and Browse
 Download
Download
 Enter Data
Enter Data
| Reaction Details | |||
|---|---|---|---|
 | Report a problem with these data | ||
| Target | Phosphatidylinositol 4,5-bisphosphate 3-kinase catalytic subunit gamma isoform | ||
| Ligand | BDBM50323729 | ||
| Substrate/Competitor | n/a | ||
| Meas. Tech. | ChEMBL_646801 (CHEMBL1216942) | ||
| IC50 | 20±n/a nM | ||
| Citation |  Berndt, A; Miller, S; Williams, O; Le, DD; Houseman, BT; Pacold, JI; Gorrec, F; Hon, WC; Liu, Y; Rommel, C; Gaillard, P; Rückle, T; Schwarz, MK; Shokat, KM; Shaw, JP; Williams, RL The p110 delta structure: mechanisms for selectivity and potency of new PI(3)K inhibitors. Nat Chem Biol6:117-24 (2010) [PubMed] Article Berndt, A; Miller, S; Williams, O; Le, DD; Houseman, BT; Pacold, JI; Gorrec, F; Hon, WC; Liu, Y; Rommel, C; Gaillard, P; Rückle, T; Schwarz, MK; Shokat, KM; Shaw, JP; Williams, RL The p110 delta structure: mechanisms for selectivity and potency of new PI(3)K inhibitors. Nat Chem Biol6:117-24 (2010) [PubMed] Article | ||
| More Info.: | Get all data from this article, Assay Method | ||
| Phosphatidylinositol 4,5-bisphosphate 3-kinase catalytic subunit gamma isoform | |||
| Name: | Phosphatidylinositol 4,5-bisphosphate 3-kinase catalytic subunit gamma isoform | ||
| Synonyms: | PI3-kinase p110 subunit gamma | PI3-kinase subunit p120-gamma | PI3Kgamma | PIK3CG | PK3CG_HUMAN | Phosphatidylinositol 4,5-biphosphate 3-kinase catalytic subunit gamma (PIK3CG) | Phosphatidylinositol 4,5-bisphosphate 3-kinase (PI3K) | Phosphatidylinositol 4,5-bisphosphate 3-kinase 110 kDa catalytic subunit gamma (PI3K gamma) | Phosphatidylinositol 4,5-bisphosphate 3-kinase catalytic subunit gamma (PI3Kgamma) | Phosphatidylinositol 4,5-bisphosphate 3-kinase catalytic subunit gamma isoform (PI3K gamma) | Phosphatidylinositol 4,5-bisphosphate 3-kinase catalytic subunit gamma isoform (PI3K) | Phosphatidylinositol 4,5-bisphosphate 3-kinase catalytic subunit gamma isoform (PI3Kgamma) | Phosphatidylinositol-4,5-bisphosphate 3-kinase catalytic subunit gamma isoform | Phosphoinositide 3-Kinase (PI3K), gamma Chain A | Phosphoinositide 3-kinases gamma (PI3K gamma) | Phosphoinositide-3-kinase (PI3K gamma) | p120-PI3K | ||
| Type: | Enzyme Subunit | ||
| Mol. Mass.: | 126470.30 | ||
| Organism: | Homo sapiens (Human) | ||
| Description: | P48736 | ||
| Residue: | 1102 | ||
| Sequence: |
| ||
| BDBM50323729 | |||
| n/a | |||
| Name | BDBM50323729 | ||
| Synonyms: | 2-((4-amino-3-(3-fluoro-4-hydroxyphenyl)-1H-pyrazolo[3,4-d]pyrimidin-1-yl)methyl)-5-methyl-3-o-tolylquinazolin-4(3H)-one | 2-{[4-amino-3-(3-fluoro-4-hydroxyphenyl)-1H-pyrazolo[3,4-d]pyrimidin-1-yl]methyl}-5-methyl-3-(2-methylphenyl)quinazolin-4(3H)-one | CHEMBL1213085 | ||
| Type | Small organic molecule | ||
| Emp. Form. | C28H22FN7O2 | ||
| Mol. Mass. | 507.5184 | ||
| SMILES | Cc1ccccc1-n1c(Cn2nc(-c3ccc(O)c(F)c3)c3c(N)ncnc23)nc2cccc(C)c2c1=O |(16.77,-15.65,;16.76,-14.1,;18.09,-13.33,;18.08,-11.78,;16.73,-11.01,;15.41,-11.8,;15.41,-13.33,;14.09,-14.11,;14.1,-15.66,;15.44,-16.43,;15.44,-17.98,;14.2,-18.89,;14.68,-20.37,;13.89,-21.68,;14.64,-23.01,;13.86,-24.33,;12.31,-24.31,;11.52,-25.62,;11.57,-22.96,;10.03,-22.93,;12.36,-21.64,;16.22,-20.35,;17.26,-21.49,;16.78,-22.96,;18.76,-21.17,;19.22,-19.7,;18.19,-18.56,;16.7,-18.89,;12.76,-16.44,;11.42,-15.67,;10.08,-16.46,;8.75,-15.68,;8.75,-14.13,;10.08,-13.36,;10.07,-11.82,;11.42,-14.12,;12.75,-13.35,;12.74,-11.81,)| | ||
| Structure | 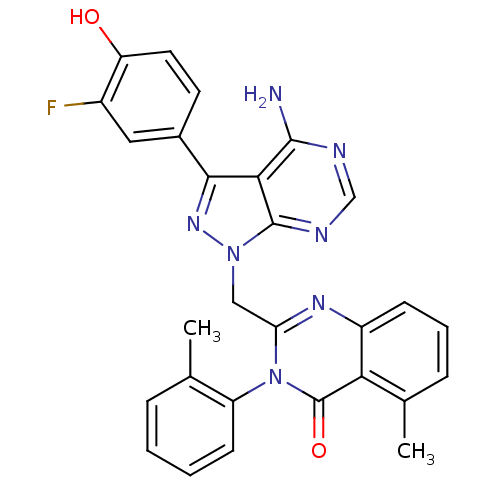
| ||