| Reaction Details |
|---|
 | Report a problem with these data |
| Target | Fatty-acid amide hydrolase 1 [30-579] |
|---|
| Ligand | BDBM26739 |
|---|
| Substrate/Competitor | n/a |
|---|
| Meas. Tech. | ChEMBL_794677 (CHEMBL1937239) |
|---|
| IC50 | 4.6±n/a nM |
|---|
| Citation |  Rusch, M; Zahov, S; Vetter, IR; Lehr, M; Hedberg, C Design, synthesis and evaluation of polar head group containing 2-keto-oxazole inhibitors of FAAH. Bioorg Med Chem20:1100-12 (2012) [PubMed] Article Rusch, M; Zahov, S; Vetter, IR; Lehr, M; Hedberg, C Design, synthesis and evaluation of polar head group containing 2-keto-oxazole inhibitors of FAAH. Bioorg Med Chem20:1100-12 (2012) [PubMed] Article |
|---|
| More Info.: | Get all data from this article, Assay Method |
|---|
| |
| Fatty-acid amide hydrolase 1 [30-579] |
|---|
| Name: | Fatty-acid amide hydrolase 1 [30-579] |
|---|
| Synonyms: | Anandamide amidohydrolase 1 | FAAH1_RAT | Faah | Faah1 | Fatty Acid Amide Hydrolase | Fatty Acid Amide Hydrolic, FAAH | Fatty-acid amide hydrolase (FAAH) | Fatty-acid amide hydrolase 1 | Fatty-acid amide hydrolase 1 (FAAH) | Fatty-acid amide hydrolase 1 (aa 30-579) | Oleamide hydrolase 1 |
|---|
| Type: | Single-pass membrane protein; homodimer |
|---|
| Mol. Mass.: | 60474.00 |
|---|
| Organism: | Rattus norvegicus (rat) |
|---|
| Description: | P97612 (aa 30-579) |
|---|
| Residue: | 550 |
|---|
| Sequence: | RWTGRQKARGAATRARQKQRASLETMDKAVQRFRLQNPDLDSEALLTLPLLQLVQKLQSG
ELSPEAVFFTYLGKAWEVNKGTNCVTSYLTDCETQLSQAPRQGLLYGVPVSLKECFSYKG
HDSTLGLSLNEGMPSESDCVVVQVLKLQGAVPFVHTNVPQSMLSFDCSNPLFGQTMNPWK
SSKSPGGSSGGEGALIGSGGSPLGLGTDIGGSIRFPSAFCGICGLKPTGNRLSKSGLKGC
VYGQTAVQLSLGPMARDVESLALCLKALLCEHLFTLDPTVPPLPFREEVYRSSRPLRVGY
YETDNYTMPSPAMRRALIETKQRLEAAGHTLIPFLPNNIPYALEVLSAGGLFSDGGRSFL
QNFKGDFVDPCLGDLILILRLPSWFKRLLSLLLKPLFPRLAAFLNSMRPRSAEKLWKLQH
EIEMYRQSVIAQWKAMNLDVLLTPMLGPALDLNTPGRATGAISYTVLYNCLDFPAGVVPV
TTVTAEDDAQMELYKGYFGDIWDIILKKAMKNSVGLPVAVQCVALPWQEELCLRFMREVE
QLMTPQKQPS
|
|
|
|---|
| BDBM26739 |
|---|
| n/a |
|---|
| Name | BDBM26739 |
|---|
| Synonyms: | 3-(3-carbamoylphenyl)phenyl N-cyclohexylcarbamate | CHEMBL184238 | URB 597 | URB-597 | URB597 | US9187413, 1a (URB597) |
|---|
| Type | Small organic molecule |
|---|
| Emp. Form. | C20H22N2O3 |
|---|
| Mol. Mass. | 338.4003 |
|---|
| SMILES | NC(=O)c1cccc(c1)-c1cccc(OC(=O)NC2CCCCC2)c1 |
|---|
| Structure | 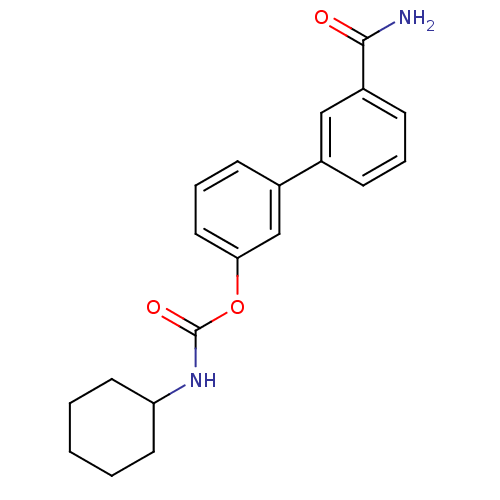
|
|---|


 Search and Browse
Search and Browse
 Download
Download
 Enter Data
Enter Data

 Rusch, M; Zahov, S; Vetter, IR; Lehr, M; Hedberg, C Design, synthesis and evaluation of polar head group containing 2-keto-oxazole inhibitors of FAAH. Bioorg Med Chem20:1100-12 (2012) [PubMed] Article
Rusch, M; Zahov, S; Vetter, IR; Lehr, M; Hedberg, C Design, synthesis and evaluation of polar head group containing 2-keto-oxazole inhibitors of FAAH. Bioorg Med Chem20:1100-12 (2012) [PubMed] Article