| Reaction Details |
|---|
 | Report a problem with these data |
| Target | Vitamin K-dependent protein C |
|---|
| Ligand | BDBM50060046 |
|---|
| Substrate/Competitor | n/a |
|---|
| Meas. Tech. | ChEMBL_27847 (CHEMBL642272) |
|---|
| Ki | 33000±n/a nM |
|---|
| Citation |  Stürzebecher, J; Prasa, D; Hauptmann, J; Vieweg, H; Wikström, P Synthesis and structure-activity relationships of potent thrombin inhibitors: piperazides of 3-amidinophenylalanine. J Med Chem40:3091-9 (1997) [PubMed] Article Stürzebecher, J; Prasa, D; Hauptmann, J; Vieweg, H; Wikström, P Synthesis and structure-activity relationships of potent thrombin inhibitors: piperazides of 3-amidinophenylalanine. J Med Chem40:3091-9 (1997) [PubMed] Article |
|---|
| More Info.: | Get all data from this article, Assay Method |
|---|
| |
| Vitamin K-dependent protein C |
|---|
| Name: | Vitamin K-dependent protein C |
|---|
| Synonyms: | Activated protein C cofactor | Anticoagulant protein C | Apolipoprotein H | Autoprothrombin IIA | Blood coagulation factor XIV | Coagulation factor V | Coagulation factor V heavy chain | Coagulation factor V light chain | Endothelial protein C receptor | PROC | PROC_HUMAN | Proaccelerin, labile factor | Vitamin K-dependent protein C precursor |
|---|
| Type: | Enzyme |
|---|
| Mol. Mass.: | 52067.73 |
|---|
| Organism: | Homo sapiens (Human) |
|---|
| Description: | n/a |
|---|
| Residue: | 461 |
|---|
| Sequence: | MWQLTSLLLFVATWGISGTPAPLDSVFSSSERAHQVLRIRKRANSFLEELRHSSLERECI
EEICDFEEAKEIFQNVDDTLAFWSKHVDGDQCLVLPLEHPCASLCCGHGTCIDGIGSFSC
DCRSGWEGRFCQREVSFLNCSLDNGGCTHYCLEEVGWRRCSCAPGYKLGDDLLQCHPAVK
FPCGRPWKRMEKKRSHLKRDTEDQEDQVDPRLIDGKMTRRGDSPWQVVLLDSKKKLACGA
VLIHPSWVLTAAHCMDESKKLLVRLGEYDLRRWEKWELDLDIKEVFVHPNYSKSTTDNDI
ALLHLAQPATLSQTIVPICLPDSGLAERELNQAGQETLVTGWGYHSSREKEAKRNRTFVL
NFIKIPVVPHNECSEVMSNMVSENMLCAGILGDRQDACEGDSGGPMVASFHGTWFLVGLV
SWGEGCGLLHNYGVYTKVSRYLDWIHGHIRDKEAPQKSWAP
|
|
|
|---|
| BDBM50060046 |
|---|
| n/a |
|---|
| Name | BDBM50060046 |
|---|
| Synonyms: | 3-[(R)-2-[2-(7,7-Dimethyl-2-oxo-bicyclo[2.2.1]hept-1-yl)-ethanesulfonylamino]-3-(4-methanesulfonyl-piperazin-1-yl)-3-oxo-propyl]-benzamidine | CHEMBL319035 |
|---|
| Type | Small organic molecule |
|---|
| Emp. Form. | C26H39N5O6S2 |
|---|
| Mol. Mass. | 581.748 |
|---|
| SMILES | CC1(C)C2CCC1(CCS(=O)(=O)N[C@H](Cc1cccc(c1)C(N)=N)C(=O)N1CCN(CC1)S(C)(=O)=O)C(=O)C2 |TLB:37:36:1:5.4| |
|---|
| Structure | 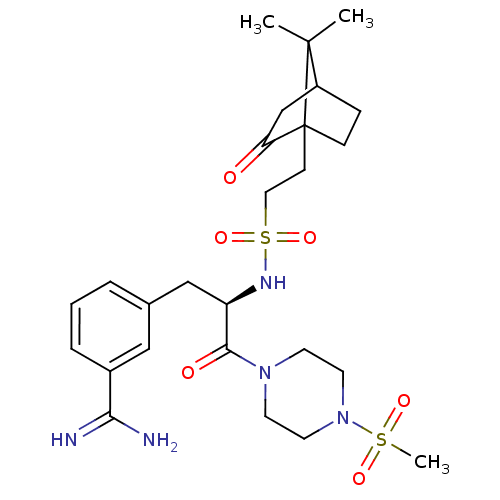
|
|---|


 Search and Browse
Search and Browse
 Download
Download
 Enter Data
Enter Data

 Stürzebecher, J; Prasa, D; Hauptmann, J; Vieweg, H; Wikström, P Synthesis and structure-activity relationships of potent thrombin inhibitors: piperazides of 3-amidinophenylalanine. J Med Chem40:3091-9 (1997) [PubMed] Article
Stürzebecher, J; Prasa, D; Hauptmann, J; Vieweg, H; Wikström, P Synthesis and structure-activity relationships of potent thrombin inhibitors: piperazides of 3-amidinophenylalanine. J Med Chem40:3091-9 (1997) [PubMed] Article