| Reaction Details |
|---|
 | Report a problem with these data |
| Target | Adenosine receptor A3 |
|---|
| Ligand | BDBM50207816 |
|---|
| Substrate/Competitor | n/a |
|---|
| Meas. Tech. | ChEMBL_31710 (CHEMBL873053) |
|---|
| Ki | 20100±n/a nM |
|---|
| Citation |  Jacobson, KA; Gao, ZG; Chen, A; Barak, D; Kim, SA; Lee, K; Link, A; Rompaey, PV; van Calenbergh, S; Liang, BT Neoceptor concept based on molecular complementarity in GPCRs: a mutant adenosine A(3) receptor with selectively enhanced affinity for amine-modified nucleosides. J Med Chem44:4125-36 (2001) [PubMed] Jacobson, KA; Gao, ZG; Chen, A; Barak, D; Kim, SA; Lee, K; Link, A; Rompaey, PV; van Calenbergh, S; Liang, BT Neoceptor concept based on molecular complementarity in GPCRs: a mutant adenosine A(3) receptor with selectively enhanced affinity for amine-modified nucleosides. J Med Chem44:4125-36 (2001) [PubMed] |
|---|
| More Info.: | Get all data from this article, Assay Method |
|---|
| |
| Adenosine receptor A3 |
|---|
| Name: | Adenosine receptor A3 |
|---|
| Synonyms: | A3 adenosine receptor (hA3) | AA3R_HUMAN | ADORA3 | Adenosine A3 receptor (A3AR) |
|---|
| Type: | G Protein-Coupled Receptor (GPCR) |
|---|
| Mol. Mass.: | 36197.32 |
|---|
| Organism: | Homo sapiens (Human) |
|---|
| Description: | P0DMS8 |
|---|
| Residue: | 318 |
|---|
| Sequence: | MPNNSTALSLANVTYITMEIFIGLCAIVGNVLVICVVKLNPSLQTTTFYFIVSLALADIA
VGVLVMPLAIVVSLGITIHFYSCLFMTCLLLIFTHASIMSLLAIAVDRYLRVKLTVRYKR
VTTHRRIWLALGLCWLVSFLVGLTPMFGWNMKLTSEYHRNVTFLSCQFVSVMRMDYMVYF
SFLTWIFIPLVVMCAIYLDIFYIIRNKLSLNLSNSKETGAFYGREFKTAKSLFLVLFLFA
LSWLPLSIINCIIYFNGEVPQLVLYMGILLSHANSMMNPIVYAYKIKKFKETYLLILKAC
VVCHPSDSLDTSIEKNSE
|
|
|
|---|
| BDBM50207816 |
|---|
| n/a |
|---|
| Name | BDBM50207816 |
|---|
| Synonyms: | CHEMBL273094 | N-(2-Amino-ethyl)-2-[4-(2,6-dioxo-1,3-dipropyl-2,3,6,7-tetrahydro-1H-purin-8-yl)-phenoxy]-acetamide | N-(2-Amino-ethyl)-2-[4-(2,6-dioxo-1,3-dipropyl-2,3,6,7-tetrahydro-1H-purin-8-yl)-phenoxy]-acetamide (xanthine amine congenere, XAC) | N-(2-Amino-ethyl)-2-[4-(2,6-dioxo-1,3-dipropyl-2,3,6,7-tetrahydro-1H-purin-8-yl)-phenoxy]-acetamide(XAC) | N-(2-Amino-ethyl)-2-[4-(2,6-dioxo-1,3-dipropyl-2,3,6,9-tetrahydro-1H-purin-8-yl)-phenoxy]-acetamide | N-(2-aminoethyl)-2-(4-(2,6-dioxo-1,3-dipropyl-2,3,6,7-tetrahydro-1H-purin-8-yl)phenoxy)acetamide | XAC |
|---|
| Type | Small organic molecule |
|---|
| Emp. Form. | C21H28N6O4 |
|---|
| Mol. Mass. | 428.4848 |
|---|
| SMILES | CCCn1c2nc([nH]c2c(=O)n(CCC)c1=O)-c1ccc(OCC(=O)NCCN)cc1 |
|---|
| Structure | 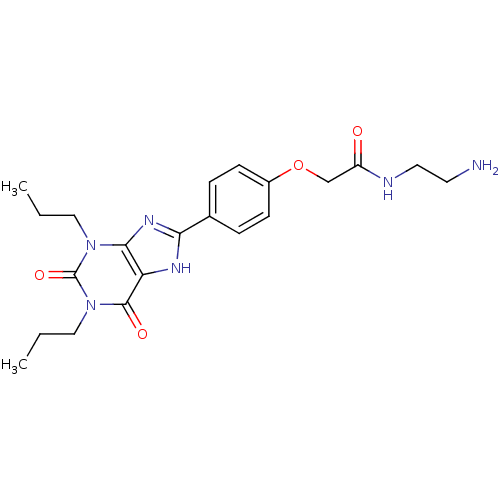
|
|---|


 Search and Browse
Search and Browse
 Download
Download
 Enter Data
Enter Data

 Jacobson, KA; Gao, ZG; Chen, A; Barak, D; Kim, SA; Lee, K; Link, A; Rompaey, PV; van Calenbergh, S; Liang, BT Neoceptor concept based on molecular complementarity in GPCRs: a mutant adenosine A(3) receptor with selectively enhanced affinity for amine-modified nucleosides. J Med Chem44:4125-36 (2001) [PubMed]
Jacobson, KA; Gao, ZG; Chen, A; Barak, D; Kim, SA; Lee, K; Link, A; Rompaey, PV; van Calenbergh, S; Liang, BT Neoceptor concept based on molecular complementarity in GPCRs: a mutant adenosine A(3) receptor with selectively enhanced affinity for amine-modified nucleosides. J Med Chem44:4125-36 (2001) [PubMed]