| Reaction Details |
|---|
 | Report a problem with these data |
| Target | Prolyl endopeptidase |
|---|
| Ligand | BDBM50170692 |
|---|
| Substrate/Competitor | n/a |
|---|
| Meas. Tech. | ChEMBL_306908 (CHEMBL828351) |
|---|
| pH | 7±n/a |
|---|
| IC50 | 1.3±n/a nM |
|---|
| Comments | extracted |
|---|
| Citation |  Jarho, EM; Wallén, EA; Christiaans, JA; Forsberg, MM; Venäläinen, JI; Männistö, PT; Gynther, J; Poso, A Dicarboxylic acid azacycle l-prolyl-pyrrolidine amides as prolyl oligopeptidase inhibitors and three-dimensional quantitative structure-activity relationship of the enzyme-inhibitor interactions. J Med Chem48:4772-82 (2005) [PubMed] Article Jarho, EM; Wallén, EA; Christiaans, JA; Forsberg, MM; Venäläinen, JI; Männistö, PT; Gynther, J; Poso, A Dicarboxylic acid azacycle l-prolyl-pyrrolidine amides as prolyl oligopeptidase inhibitors and three-dimensional quantitative structure-activity relationship of the enzyme-inhibitor interactions. J Med Chem48:4772-82 (2005) [PubMed] Article |
|---|
| More Info.: | Get all data from this article, Assay Method |
|---|
| |
| Prolyl endopeptidase |
|---|
| Name: | Prolyl endopeptidase |
|---|
| Synonyms: | 3.4.21.26 | PE | PPCE_PIG | PREP | Post-proline cleaving enzyme |
|---|
| Type: | n/a |
|---|
| Mol. Mass.: | 80758.04 |
|---|
| Organism: | Sus scrofa |
|---|
| Description: | n/a |
|---|
| Residue: | 710 |
|---|
| Sequence: | MLSFQYPDVYRDETAIQDYHGHKVCDPYAWLEDPDSEQTKAFVEAQNKITVPFLEQCPIR
GLYKERMTELYDYPKYSCHFKKGKRYFYFYNTGLQNQRVLYVQDSLEGEARVFLDPNILS
DDGTVALRGYAFSEDGEYFAYGLSASGSDWVTIKFMKVDGAKELPDVLERVKFSCMAWTH
DGKGMFYNAYPQQDGKSDGTETSTNLHQKLYYHVLGTDQSEDILCAEFPDEPKWMGGAEL
SDDGRYVLLSIREGCDPVNRLWYCDLQQESNGITGILKWVKLIDNFEGEYDYVTNEGTVF
TFKTNRHSPNYRLINIDFTDPEESKWKVLVPEHEKDVLEWVACVRSNFLVLCYLHDVKNT
LQLHDLATGALLKIFPLEVGSVVGYSGQKKDTEIFYQFTSFLSPGIIYHCDLTKEELEPR
VFREVTVKGIDASDYQTVQIFYPSKDGTKIPMFIVHKKGIKLDGSHPAFLYGYGGFNISI
TPNYSVSRLIFVRHMGGVLAVANIRGGGEYGETWHKGGILANKQNCFDDFQCAAEYLIKE
GYTSPKRLTINGGSNGGLLVATCANQRPDLFGCVIAQVGVMDMLKFHKYTIGHAWTTDYG
CSDSKQHFEWLIKYSPLHNVKLPEADDIQYPSMLLLTADHDDRVVPLHSLKFIATLQYIV
GRSRKQNNPLLIHVDTKAGHGAGKPTAKVIEEVSDMFAFIARCLNIDWIP
|
|
|
|---|
| BDBM50170692 |
|---|
| n/a |
|---|
| Name | BDBM50170692 |
|---|
| Synonyms: | (S)-1-{(S)-1-[3-((S)-2-Benzoyl-pyrrolidine-1-carbonyl)-benzoyl]-pyrrolidine-2-carbonyl}-pyrrolidine-2-carbonitrile | CHEMBL364989 |
|---|
| Type | Small organic molecule |
|---|
| Emp. Form. | C29H30N4O4 |
|---|
| Mol. Mass. | 498.5729 |
|---|
| SMILES | O=C([C@@H]1CCCN1C(=O)c1cccc(c1)C(=O)N1CCC[C@H]1C(=O)c1ccccc1)N1CCC[C@H]1C#N |
|---|
| Structure | 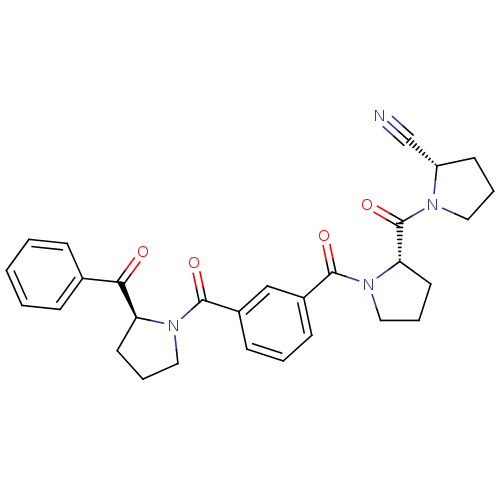
|
|---|


 Search and Browse
Search and Browse
 Download
Download
 Enter Data
Enter Data

 Jarho, EM; Wallén, EA; Christiaans, JA; Forsberg, MM; Venäläinen, JI; Männistö, PT; Gynther, J; Poso, A Dicarboxylic acid azacycle l-prolyl-pyrrolidine amides as prolyl oligopeptidase inhibitors and three-dimensional quantitative structure-activity relationship of the enzyme-inhibitor interactions. J Med Chem48:4772-82 (2005) [PubMed] Article
Jarho, EM; Wallén, EA; Christiaans, JA; Forsberg, MM; Venäläinen, JI; Männistö, PT; Gynther, J; Poso, A Dicarboxylic acid azacycle l-prolyl-pyrrolidine amides as prolyl oligopeptidase inhibitors and three-dimensional quantitative structure-activity relationship of the enzyme-inhibitor interactions. J Med Chem48:4772-82 (2005) [PubMed] Article