| Reaction Details |
|---|
 | Report a problem with these data |
| Target | Heat shock protein HSP 90-alpha |
|---|
| Ligand | BDBM20926 |
|---|
| Substrate/Competitor | n/a |
|---|
| Meas. Tech. | ChEMBL_616197 (CHEMBL1100282) |
|---|
| Ki | 9±n/a nM |
|---|
| Citation |  Biamonte, MA; Van de Water, R; Arndt, JW; Scannevin, RH; Perret, D; Lee, WC Heat shock protein 90: inhibitors in clinical trials. J Med Chem53:3-17 (2010) [PubMed] Article Biamonte, MA; Van de Water, R; Arndt, JW; Scannevin, RH; Perret, D; Lee, WC Heat shock protein 90: inhibitors in clinical trials. J Med Chem53:3-17 (2010) [PubMed] Article |
|---|
| More Info.: | Get all data from this article, Assay Method |
|---|
| |
| Heat shock protein HSP 90-alpha |
|---|
| Name: | Heat shock protein HSP 90-alpha |
|---|
| Synonyms: | HS90A_HUMAN | HSP 86 | HSP86 | HSP90A | HSP90AA1 | HSPC1 | HSPCA | Heat Shock Protein 90 (Hsp90) | Heat shock 86 kDa | Heat shock protein HSP 90 (HSP90) | Heat shock protein HSP 90-alpha (HSP90) | Heat shock protein HSP 90-alpha (HSP90A) | LAP-2 | LPS-associated protein 2 | Lipopolysaccharide-associated protein 2 | Renal carcinoma antigen NY-REN-38 | heat shock protein 90kDa alpha (cytosolic), class A member 1 isoform 2 |
|---|
| Type: | Molecular Chaperone |
|---|
| Mol. Mass.: | 84623.45 |
|---|
| Organism: | Homo sapiens (Human) |
|---|
| Description: | P07900 |
|---|
| Residue: | 732 |
|---|
| Sequence: | MPEETQTQDQPMEEEEVETFAFQAEIAQLMSLIINTFYSNKEIFLRELISNSSDALDKIR
YESLTDPSKLDSGKELHINLIPNKQDRTLTIVDTGIGMTKADLINNLGTIAKSGTKAFME
ALQAGADISMIGQFGVGFYSAYLVAEKVTVITKHNDDEQYAWESSAGGSFTVRTDTGEPM
GRGTKVILHLKEDQTEYLEERRIKEIVKKHSQFIGYPITLFVEKERDKEVSDDEAEEKED
KEEEKEKEEKESEDKPEIEDVGSDEEEEKKDGDKKKKKKIKEKYIDQEELNKTKPIWTRN
PDDITNEEYGEFYKSLTNDWEDHLAVKHFSVEGQLEFRALLFVPRRAPFDLFENRKKKNN
IKLYVRRVFIMDNCEELIPEYLNFIRGVVDSEDLPLNISREMLQQSKILKVIRKNLVKKC
LELFTELAEDKENYKKFYEQFSKNIKLGIHEDSQNRKKLSELLRYYTSASGDEMVSLKDY
CTRMKENQKHIYYITGETKDQVANSAFVERLRKHGLEVIYMIEPIDEYCVQQLKEFEGKT
LVSVTKEGLELPEDEEEKKKQEEKKTKFENLCKIMKDILEKKVEKVVVSNRLVTSPCCIV
TSTYGWTANMERIMKAQALRDNSTMGYMAAKKHLEINPDHSIIETLRQKAEADKNDKSVK
DLVILLYETALLSSGFSLEDPQTHANRIYRMIKLGLGIDEDDPTADDTSAAVTEEMPPLE
GDDDTSRMEEVD
|
|
|
|---|
| BDBM20926 |
|---|
| n/a |
|---|
| Name | BDBM20926 |
|---|
| Synonyms: | 5-[2,4-dihydroxy-5-(propan-2-yl)phenyl]-N-ethyl-4-[4-(morpholin-4-ylmethyl)phenyl]-1,2-oxazole-3-carboxamide | Isoxazole, 40f | VER-52296/NVP-AUY922 |
|---|
| Type | Small organic molecule |
|---|
| Emp. Form. | C26H31N3O5 |
|---|
| Mol. Mass. | 465.5414 |
|---|
| SMILES | CCNC(=O)c1noc(c1-c1ccc(CN2CCOCC2)cc1)-c1cc(C(C)C)c(O)cc1O |
|---|
| Structure | 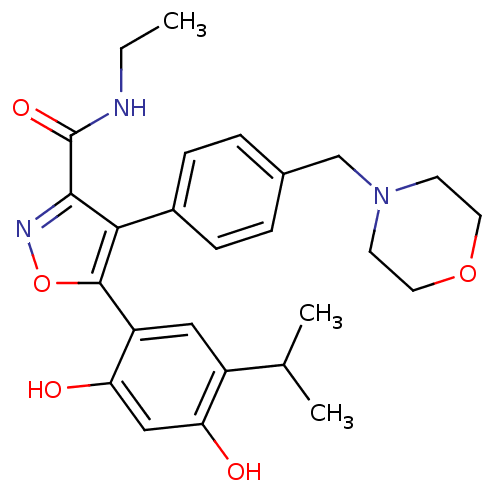
|
|---|


 Search and Browse
Search and Browse
 Download
Download
 Enter Data
Enter Data

 Biamonte, MA; Van de Water, R; Arndt, JW; Scannevin, RH; Perret, D; Lee, WC Heat shock protein 90: inhibitors in clinical trials. J Med Chem53:3-17 (2010) [PubMed] Article
Biamonte, MA; Van de Water, R; Arndt, JW; Scannevin, RH; Perret, D; Lee, WC Heat shock protein 90: inhibitors in clinical trials. J Med Chem53:3-17 (2010) [PubMed] Article