| Reaction Details |
|---|
 | Report a problem with these data |
| Target | cGMP-specific 3',5'-cyclic phosphodiesterase |
|---|
| Ligand | BDBM50316638 |
|---|
| Substrate/Competitor | n/a |
|---|
| Meas. Tech. | ChEMBL_632193 (CHEMBL1106526) |
|---|
| IC50 | 0.01±n/a nM |
|---|
| Citation |  Tollefson, MB; Acker, BA; Jacobsen, EJ; Hughes, RO; Walker, JK; Fox, DN; Palmer, MJ; Freeman, SK; Yu, Y; Bond, BR 1-(2-Ethoxyethyl)-1H-pyrazolo[4,3-d]pyrimidines as potent phosphodiesterase 5 (PDE5) inhibitors. Bioorg Med Chem Lett20:3120-4 (2010) [PubMed] Tollefson, MB; Acker, BA; Jacobsen, EJ; Hughes, RO; Walker, JK; Fox, DN; Palmer, MJ; Freeman, SK; Yu, Y; Bond, BR 1-(2-Ethoxyethyl)-1H-pyrazolo[4,3-d]pyrimidines as potent phosphodiesterase 5 (PDE5) inhibitors. Bioorg Med Chem Lett20:3120-4 (2010) [PubMed] |
|---|
| More Info.: | Get all data from this article, Assay Method |
|---|
| |
| cGMP-specific 3',5'-cyclic phosphodiesterase |
|---|
| Name: | cGMP-specific 3',5'-cyclic phosphodiesterase |
|---|
| Synonyms: | 3',5'-cyclic phosphodiesterase | CGB-PDE | PDE5 | PDE5A | PDE5A_HUMAN | Phosphodiesterase 2 and 5 (PDE2 and PDE5) | Phosphodiesterase 5 (PDE5) | Phosphodiesterase 5A | Phosphodiesterase 5A (PDE5A) | cGMP-binding cGMP-specific phosphodiesterase | cGMP-specific 3',5'-cyclic phosphodiesterase |
|---|
| Type: | Protein |
|---|
| Mol. Mass.: | 99975.83 |
|---|
| Organism: | Homo sapiens (Human) |
|---|
| Description: | O76074 |
|---|
| Residue: | 875 |
|---|
| Sequence: | MERAGPSFGQQRQQQQPQQQKQQQRDQDSVEAWLDDHWDFTFSYFVRKATREMVNAWFAE
RVHTIPVCKEGIRGHTESCSCPLQQSPRADNSAPGTPTRKISASEFDRPLRPIVVKDSEG
TVSFLSDSEKKEQMPLTPPRFDHDEGDQCSRLLELVKDISSHLDVTALCHKIFLHIHGLI
SADRYSLFLVCEDSSNDKFLISRLFDVAEGSTLEEVSNNCIRLEWNKGIVGHVAALGEPL
NIKDAYEDPRFNAEVDQITGYKTQSILCMPIKNHREEVVGVAQAINKKSGNGGTFTEKDE
KDFAAYLAFCGIVLHNAQLYETSLLENKRNQVLLDLASLIFEEQQSLEVILKKIAATIIS
FMQVQKCTIFIVDEDCSDSFSSVFHMECEELEKSSDTLTREHDANKINYMYAQYVKNTME
PLNIPDVSKDKRFPWTTENTGNVNQQCIRSLLCTPIKNGKKNKVIGVCQLVNKMEENTGK
VKPFNRNDEQFLEAFVIFCGLGIQNTQMYEAVERAMAKQMVTLEVLSYHASAAEEETREL
QSLAAAVVPSAQTLKITDFSFSDFELSDLETALCTIRMFTDLNLVQNFQMKHEVLCRWIL
SVKKNYRKNVAYHNWRHAFNTAQCMFAALKAGKIQNKLTDLEILALLIAALSHDLDHRGV
NNSYIQRSEHPLAQLYCHSIMEHHHFDQCLMILNSPGNQILSGLSIEEYKTTLKIIKQAI
LATDLALYIKRRGEFFELIRKNQFNLEDPHQKELFLAMLMTACDLSAITKPWPIQQRIAE
LVATEFFDQGDRERKELNIEPTDLMNREKKNKIPSMQVGFIDAICLQLYEALTHVSEDCF
PLLDGCRKNRQKWQALAEQQEKMLINGESGQAKRN
|
|
|
|---|
| BDBM50316638 |
|---|
| n/a |
|---|
| Name | BDBM50316638 |
|---|
| Synonyms: | 1-(2-ethoxyethyl)-3-ethyl-N7-(4-methylpyridin-2-yl)-N5-(piperidin-4-ylmethyl)-1H-pyrazolo[4,3-d]pyrimidine-5,7-diamine | CHEMBL1098465 |
|---|
| Type | Small organic molecule |
|---|
| Emp. Form. | C23H34N8O |
|---|
| Mol. Mass. | 438.5691 |
|---|
| SMILES | CCOCCn1nc(CC)c2nc(NCC3CCNCC3)nc(Nc3cc(C)ccn3)c12 |
|---|
| Structure | 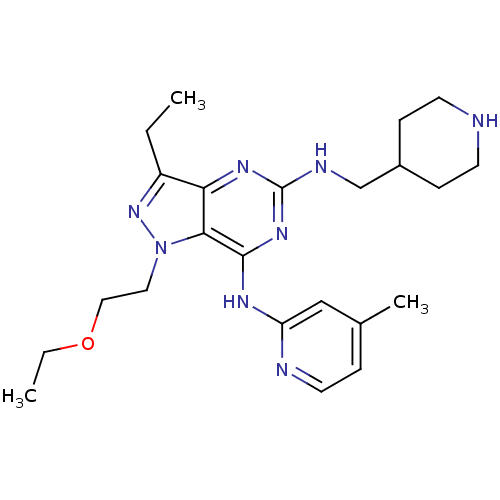
|
|---|


 Search and Browse
Search and Browse
 Download
Download
 Enter Data
Enter Data

 Tollefson, MB; Acker, BA; Jacobsen, EJ; Hughes, RO; Walker, JK; Fox, DN; Palmer, MJ; Freeman, SK; Yu, Y; Bond, BR 1-(2-Ethoxyethyl)-1H-pyrazolo[4,3-d]pyrimidines as potent phosphodiesterase 5 (PDE5) inhibitors. Bioorg Med Chem Lett20:3120-4 (2010) [PubMed]
Tollefson, MB; Acker, BA; Jacobsen, EJ; Hughes, RO; Walker, JK; Fox, DN; Palmer, MJ; Freeman, SK; Yu, Y; Bond, BR 1-(2-Ethoxyethyl)-1H-pyrazolo[4,3-d]pyrimidines as potent phosphodiesterase 5 (PDE5) inhibitors. Bioorg Med Chem Lett20:3120-4 (2010) [PubMed]