| Reaction Details |
|---|
 | Report a problem with these data |
| Target | Poly [ADP-ribose] polymerase tankyrase-1 |
|---|
| Ligand | BDBM50294835 |
|---|
| Substrate/Competitor | n/a |
|---|
| Meas. Tech. | ChEMBL_818394 (CHEMBL2034621) |
|---|
| IC50 | 131±n/a nM |
|---|
| Citation |  Narwal, M; Venkannagari, H; Lehtiö, L Structural basis of selective inhibition of human tankyrases. J Med Chem55:1360-7 (2012) [PubMed] Article Narwal, M; Venkannagari, H; Lehtiö, L Structural basis of selective inhibition of human tankyrases. J Med Chem55:1360-7 (2012) [PubMed] Article |
|---|
| More Info.: | Get all data from this article, Assay Method |
|---|
| |
| Poly [ADP-ribose] polymerase tankyrase-1 |
|---|
| Name: | Poly [ADP-ribose] polymerase tankyrase-1 |
|---|
| Synonyms: | (ARTD5 or PARP5a) | PARP5A | PARPL | Poly [ADP-ribose] polymerase 5 (PARP5) | Poly [ADP-ribose] polymerase tankyrase-1 | TIN1 | TINF1 | TNKS | TNKS1 | TNKS1_HUMAN | Tankyrase 1 | Tankyrase 1/2 | Tankyrase-1 | Tankyrase-1 (TNKS-1) | Tankyrase-1 (TNKS1) |
|---|
| Type: | Enzyme |
|---|
| Mol. Mass.: | 142058.03 |
|---|
| Organism: | Homo sapiens (Human) |
|---|
| Description: | O95271 |
|---|
| Residue: | 1327 |
|---|
| Sequence: | MAASRRSQHHHHHHQQQLQPAPGASAPPPPPPPPLSPGLAPGTTPASPTASGLAPFASPR
HGLALPEGDGSRDPPDRPRSPDPVDGTSCCSTTSTICTVAAAPVVPAVSTSSAAGVAPNP
AGSGSNNSPSSSSSPTSSSSSSPSSPGSSLAESPEAAGVSSTAPLGPGAAGPGTGVPAVS
GALRELLEACRNGDVSRVKRLVDAANVNAKDMAGRKSSPLHFAAGFGRKDVVEHLLQMGA
NVHARDDGGLIPLHNACSFGHAEVVSLLLCQGADPNARDNWNYTPLHEAAIKGKIDVCIV
LLQHGADPNIRNTDGKSALDLADPSAKAVLTGEYKKDELLEAARSGNEEKLMALLTPLNV
NCHASDGRKSTPLHLAAGYNRVRIVQLLLQHGADVHAKDKGGLVPLHNACSYGHYEVTEL
LLKHGACVNAMDLWQFTPLHEAASKNRVEVCSLLLSHGADPTLVNCHGKSAVDMAPTPEL
RERLTYEFKGHSLLQAAREADLAKVKKTLALEIINFKQPQSHETALHCAVASLHPKRKQV
TELLLRKGANVNEKNKDFMTPLHVAAERAHNDVMEVLHKHGAKMNALDTLGQTALHRAAL
AGHLQTCRLLLSYGSDPSIISLQGFTAAQMGNEAVQQILSESTPIRTSDVDYRLLEASKA
GDLETVKQLCSSQNVNCRDLEGRHSTPLHFAAGYNRVSVVEYLLHHGADVHAKDKGGLVP
LHNACSYGHYEVAELLVRHGASVNVADLWKFTPLHEAAAKGKYEICKLLLKHGADPTKKN
RDGNTPLDLVKEGDTDIQDLLRGDAALLDAAKKGCLARVQKLCTPENINCRDTQGRNSTP
LHLAAGYNNLEVAEYLLEHGADVNAQDKGGLIPLHNAASYGHVDIAALLIKYNTCVNATD
KWAFTPLHEAAQKGRTQLCALLLAHGADPTMKNQEGQTPLDLATADDIRALLIDAMPPEA
LPTCFKPQATVVSASLISPASTPSCLSAASSIDNLTGPLAELAVGGASNAGDGAAGTERK
EGEVAGLDMNISQFLKSLGLEHLRDIFETEQITLDVLADMGHEELKEIGINAYGHRHKLI
KGVERLLGGQQGTNPYLTFHCVNQGTILLDLAPEDKEYQSVEEEMQSTIREHRDGGNAGG
IFNRYNVIRIQKVVNKKLRERFCHRQKEVSEENHNHHNERMLFHGSPFINAIIHKGFDER
HAYIGGMFGAGIYFAENSSKSNQYVYGIGGGTGCPTHKDRSCYICHRQMLFCRVTLGKSF
LQFSTMKMAHAPPGHHSVIGRPSVNGLAYAEYVIYRGEQAYPEYLITYQIMKPEAPSQTA
TAAEQKT
|
|
|
|---|
| BDBM50294835 |
|---|
| n/a |
|---|
| Name | BDBM50294835 |
|---|
| Synonyms: | 4-((1S,2S,6R,7R)-3,5-Dioxo-4-aza-tricyclo[5.2.1.0*2,6*]dec-8-en-4-yl)-N-methyl-N-quinolin-8-yl-benzamide | CHEMBL551505 | CHEMBL562310 | N-(Quinolin-8-yl)-4-(endo-4-aza-3,5-dioxotricyclo[5.2.1.02,6]oct-8-en-4-yl)benzamide, 1 |
|---|
| Type | Small organic molecule |
|---|
| Emp. Form. | C25H19N3O3 |
|---|
| Mol. Mass. | 409.4367 |
|---|
| SMILES | Oc1c2[C@H]3C[C@H](C=C3)c2c(O)n1-c1ccc(cc1)C(=O)Nc1cccc2cccnc12 |r,c:6| |
|---|
| Structure | 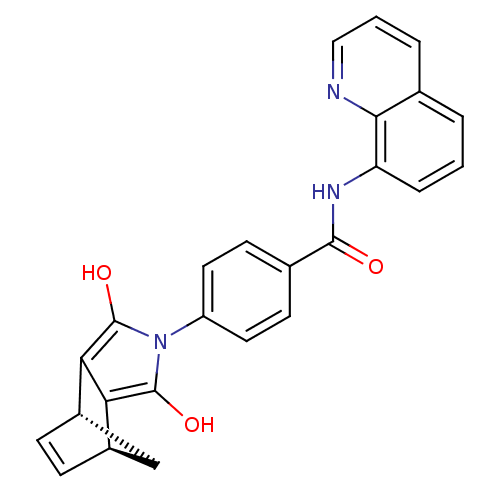
|
|---|


 Search and Browse
Search and Browse
 Download
Download
 Enter Data
Enter Data

 Narwal, M; Venkannagari, H; Lehtiö, L Structural basis of selective inhibition of human tankyrases. J Med Chem55:1360-7 (2012) [PubMed] Article
Narwal, M; Venkannagari, H; Lehtiö, L Structural basis of selective inhibition of human tankyrases. J Med Chem55:1360-7 (2012) [PubMed] Article