| Reaction Details |
|---|
 | Report a problem with these data |
| Target | P2X purinoceptor 7 |
|---|
| Ligand | BDBM50411901 |
|---|
| Substrate/Competitor | n/a |
|---|
| Meas. Tech. | ChEMBL_469375 (CHEMBL932852) |
|---|
| IC50 | 67.61±n/a nM |
|---|
| Citation |  Florjancic, AS; Peddi, S; Perez-Medrano, A; Li, B; Namovic, MT; Grayson, G; Donnelly-Roberts, DL; Jarvis, MF; Carroll, WA Synthesis and in vitro activity of 1-(2,3-dichlorophenyl)-N-(pyridin-3-ylmethyl)-1H-1,2,4-triazol-5-amine and 4-(2,3-dichlorophenyl)-N-(pyridin-3-ylmethyl)-4H-1,2,4-triazol-3-amine P2X7 antagonists. Bioorg Med Chem Lett18:2089-92 (2008) [PubMed] Article Florjancic, AS; Peddi, S; Perez-Medrano, A; Li, B; Namovic, MT; Grayson, G; Donnelly-Roberts, DL; Jarvis, MF; Carroll, WA Synthesis and in vitro activity of 1-(2,3-dichlorophenyl)-N-(pyridin-3-ylmethyl)-1H-1,2,4-triazol-5-amine and 4-(2,3-dichlorophenyl)-N-(pyridin-3-ylmethyl)-4H-1,2,4-triazol-3-amine P2X7 antagonists. Bioorg Med Chem Lett18:2089-92 (2008) [PubMed] Article |
|---|
| More Info.: | Get all data from this article, Assay Method |
|---|
| |
| P2X purinoceptor 7 |
|---|
| Name: | P2X purinoceptor 7 |
|---|
| Synonyms: | ATP receptor | P2RX7 | P2RX7_HUMAN | P2X purinoceptor 7 (P2RX7) | P2X purinoceptor 7 (P2X7) | P2X7 | P2Z receptor | Purinergic receptor |
|---|
| Type: | Protein |
|---|
| Mol. Mass.: | 68602.85 |
|---|
| Organism: | Homo sapiens (Human) |
|---|
| Description: | Q99572 |
|---|
| Residue: | 595 |
|---|
| Sequence: | MPACCSCSDVFQYETNKVTRIQSMNYGTIKWFFHVIIFSYVCFALVSDKLYQRKEPVISS
VHTKVKGIAEVKEEIVENGVKKLVHSVFDTADYTFPLQGNSFFVMTNFLKTEGQEQRLCP
EYPTRRTLCSSDRGCKKGWMDPQSKGIQTGRCVVYEGNQKTCEVSAWCPIEAVEEAPRPA
LLNSAENFTVLIKNNIDFPGHNYTTRNILPGLNITCTFHKTQNPQCPIFRLGDIFRETGD
NFSDVAIQGGIMGIEIYWDCNLDRWFHHCRPKYSFRRLDDKTTNVSLYPGYNFRYAKYYK
ENNVEKRTLIKVFGIRFDILVFGTGGKFDIIQLVVYIGSTLSYFGLAAVFIDFLIDTYSS
NCCRSHIYPWCKCCQPCVVNEYYYRKKCESIVEPKPTLKYVSFVDESHIRMVNQQLLGRS
LQDVKGQEVPRPAMDFTDLSRLPLALHDTPPIPGQPEEIQLLRKEATPRSRDSPVWCQCG
SCLPSQLPESHRCLEELCCRKKPGACITTSELFRKLVLSRHVLQFLLLYQEPLLALDVDS
TNSRLRHCAYRCYATWRFGSQDMADFAILPSCCRWRIRKEFPKSEGQYSGFKSPY
|
|
|
|---|
| BDBM50411901 |
|---|
| n/a |
|---|
| Name | BDBM50411901 |
|---|
| Synonyms: | CHEMBL270436 |
|---|
| Type | Small organic molecule |
|---|
| Emp. Form. | C22H27Cl2N7 |
|---|
| Mol. Mass. | 460.403 |
|---|
| SMILES | CC(C)N1CCCN(CC1)c1ncccc1CNc1nncn1-c1cccc(Cl)c1Cl |
|---|
| Structure | 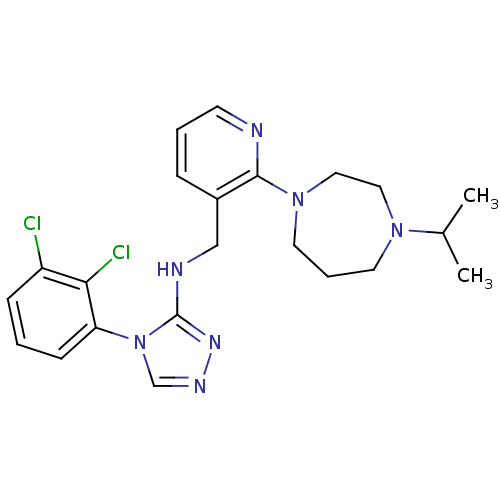
|
|---|


 Search and Browse
Search and Browse
 Download
Download
 Enter Data
Enter Data

 Florjancic, AS; Peddi, S; Perez-Medrano, A; Li, B; Namovic, MT; Grayson, G; Donnelly-Roberts, DL; Jarvis, MF; Carroll, WA Synthesis and in vitro activity of 1-(2,3-dichlorophenyl)-N-(pyridin-3-ylmethyl)-1H-1,2,4-triazol-5-amine and 4-(2,3-dichlorophenyl)-N-(pyridin-3-ylmethyl)-4H-1,2,4-triazol-3-amine P2X7 antagonists. Bioorg Med Chem Lett18:2089-92 (2008) [PubMed] Article
Florjancic, AS; Peddi, S; Perez-Medrano, A; Li, B; Namovic, MT; Grayson, G; Donnelly-Roberts, DL; Jarvis, MF; Carroll, WA Synthesis and in vitro activity of 1-(2,3-dichlorophenyl)-N-(pyridin-3-ylmethyl)-1H-1,2,4-triazol-5-amine and 4-(2,3-dichlorophenyl)-N-(pyridin-3-ylmethyl)-4H-1,2,4-triazol-3-amine P2X7 antagonists. Bioorg Med Chem Lett18:2089-92 (2008) [PubMed] Article