| Reaction Details |
|---|
 | Report a problem with these data |
| Target | Nociceptin receptor |
|---|
| Ligand | BDBM50333104 |
|---|
| Substrate/Competitor | n/a |
|---|
| Meas. Tech. | ChEMBL_964181 (CHEMBL2394994) |
|---|
| IC50 | 0.290000±n/a nM |
|---|
| Citation |  Veinberg, G; Vorona, M; Zvejniece, L; Vilskersts, R; Vavers, E; Liepinsh, E; Kazoka, H; Belyakov, S; Mishnev, A; Kuznecovs, J; Vikainis, S; Orlova, N; Lebedev, A; Ponomaryov, Y; Dambrova, M Synthesis and biological evaluation of 2-(5-methyl-4-phenyl-2-oxopyrrolidin-1-yl)-acetamide stereoisomers as novel positive allosteric modulators of sigma-1 receptor. Bioorg Med Chem21:2764-71 (2013) [PubMed] Article Veinberg, G; Vorona, M; Zvejniece, L; Vilskersts, R; Vavers, E; Liepinsh, E; Kazoka, H; Belyakov, S; Mishnev, A; Kuznecovs, J; Vikainis, S; Orlova, N; Lebedev, A; Ponomaryov, Y; Dambrova, M Synthesis and biological evaluation of 2-(5-methyl-4-phenyl-2-oxopyrrolidin-1-yl)-acetamide stereoisomers as novel positive allosteric modulators of sigma-1 receptor. Bioorg Med Chem21:2764-71 (2013) [PubMed] Article |
|---|
| More Info.: | Get all data from this article, Assay Method |
|---|
| |
| Nociceptin receptor |
|---|
| Name: | Nociceptin receptor |
|---|
| Synonyms: | KOR-3 | Kappa-type 3 opioid receptor | Mu-type opioid receptor (Mu) | NOP | Nociceptin Receptor (ORL1 Receptor) | Nociceptin receptor (NOP) | Nociceptin receptor (ORL-1) | Nociceptin receptor (ORL1) | Nociceptin/Orphanin FQ, NOP receptor | OOR | OPIATE ORL-1 | OPRL1 | OPRL1 protein | OPRX_HUMAN | ORL1 | ORL1 receptor | Opioid receptor like-1 | Orphanin FQ receptor | Orphanin FQ receptor (ORL1) | P41146 |
|---|
| Type: | G Protein-Coupled Receptor (GPCR) |
|---|
| Mol. Mass.: | 40702.87 |
|---|
| Organism: | Homo sapiens (Human) |
|---|
| Description: | P41146 |
|---|
| Residue: | 370 |
|---|
| Sequence: | MEPLFPAPFWEVIYGSHLQGNLSLLSPNHSLLPPHLLLNASHGAFLPLGLKVTIVGLYLA
VCVGGLLGNCLVMYVILRHTKMKTATNIYIFNLALADTLVLLTLPFQGTDILLGFWPFGN
ALCKTVIAIDYYNMFTSTFTLTAMSVDRYVAICHPIRALDVRTSSKAQAVNVAIWALASV
VGVPVAIMGSAQVEDEEIECLVEIPTPQDYWGPVFAICIFLFSFIVPVLVISVCYSLMIR
RLRGVRLLSGSREKDRNLRRITRLVLVVVAVFVGCWTPVQVFVLAQGLGVQPSSETAVAI
LRFCTALGYVNSCLNPILYAFLDENFKACFRKFCCASALRRDVQVSDRVRSIAKDVALAC
KTSETVPRPA
|
|
|
|---|
| BDBM50333104 |
|---|
| n/a |
|---|
| Name | BDBM50333104 |
|---|
| Synonyms: | CHEMBL389521 | H-FGGFTGARKSARKLANQ-NH2 | N/OFQ-NH2 | NC(1-17)NH2 | Nociceptin-NH2 | Nociceptin/orphanin FQ |
|---|
| Type | Small organic molecule |
|---|
| Emp. Form. | C79H130N28O21 |
|---|
| Mol. Mass. | 1808.0525 |
|---|
| SMILES | [#6]-[#6](-[#6])-[#6]-[#6@H](-[#7]-[#6](=O)-[#6@H](-[#6]-[#6]-[#6]-[#6]-[#7])-[#7]-[#6](=O)-[#6@H](-[#6]-[#6]-[#6]\[#7]=[#6](/[#7])-[#7])-[#7]-[#6](=O)-[#6@H](-[#6])-[#7]-[#6](=O)-[#6@H](-[#6]-[#8])-[#7]-[#6](=O)-[#6@H](-[#6]-[#6]-[#6]-[#6]-[#7])-[#7]-[#6](=O)-[#6@H](-[#6]-[#6]-[#6]\[#7]=[#6](/[#7])-[#7])-[#7]-[#6](=O)-[#6@H](-[#6])-[#7]-[#6](=O)-[#6]-[#7]-[#6](=O)-[#6@@H](-[#7]-[#6](=O)-[#6@H](-[#6]-c1ccccc1)-[#7]-[#6](=O)-[#6]-[#7]-[#6](=O)-[#6]-[#7]-[#6](=O)-[#6@@H](-[#7])-[#6]-c1ccccc1)-[#6@@H](-[#6])-[#8])-[#6](=O)-[#7]-[#6@@H](-[#6])-[#6](=O)-[#7]-[#6@@H](-[#6]-[#6](-[#7])=O)-[#6](=O)-[#7]-[#6@@H](-[#6]-[#6]-[#6](-[#7])=O)-[#6](-[#7])=O |
|---|
| Structure | 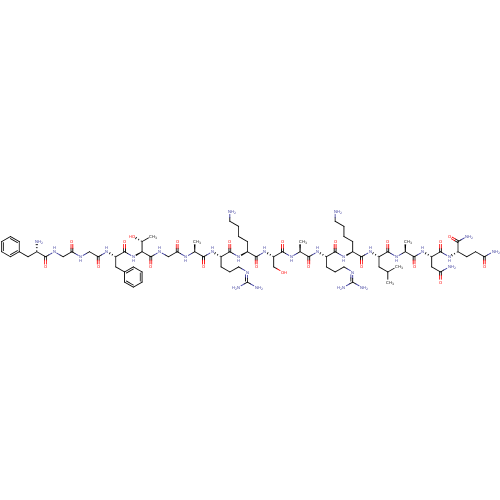
|
|---|


 Search and Browse
Search and Browse
 Download
Download
 Enter Data
Enter Data

 Veinberg, G; Vorona, M; Zvejniece, L; Vilskersts, R; Vavers, E; Liepinsh, E; Kazoka, H; Belyakov, S; Mishnev, A; Kuznecovs, J; Vikainis, S; Orlova, N; Lebedev, A; Ponomaryov, Y; Dambrova, M Synthesis and biological evaluation of 2-(5-methyl-4-phenyl-2-oxopyrrolidin-1-yl)-acetamide stereoisomers as novel positive allosteric modulators of sigma-1 receptor. Bioorg Med Chem21:2764-71 (2013) [PubMed] Article
Veinberg, G; Vorona, M; Zvejniece, L; Vilskersts, R; Vavers, E; Liepinsh, E; Kazoka, H; Belyakov, S; Mishnev, A; Kuznecovs, J; Vikainis, S; Orlova, N; Lebedev, A; Ponomaryov, Y; Dambrova, M Synthesis and biological evaluation of 2-(5-methyl-4-phenyl-2-oxopyrrolidin-1-yl)-acetamide stereoisomers as novel positive allosteric modulators of sigma-1 receptor. Bioorg Med Chem21:2764-71 (2013) [PubMed] Article