| Reaction Details |
|---|
 | Report a problem with these data |
| Target | Aminopeptidase N |
|---|
| Ligand | BDBM50024594 |
|---|
| Substrate/Competitor | n/a |
|---|
| Meas. Tech. | ChEMBL_1442055 (CHEMBL3373423) |
|---|
| Ki | 53±n/a nM |
|---|
| Citation |  Vassiliou, S; Weglarz-Tomczak, E; Berlicki, L; Pawelczak, M; Nocek, B; Mulligan, R; Joachimiak, A; Mucha, A Structure-guided, single-point modifications in the phosphinic dipeptide structure yield highly potent and selective inhibitors of neutral aminopeptidases. J Med Chem57:8140-51 (2014) [PubMed] Article Vassiliou, S; Weglarz-Tomczak, E; Berlicki, L; Pawelczak, M; Nocek, B; Mulligan, R; Joachimiak, A; Mucha, A Structure-guided, single-point modifications in the phosphinic dipeptide structure yield highly potent and selective inhibitors of neutral aminopeptidases. J Med Chem57:8140-51 (2014) [PubMed] Article |
|---|
| More Info.: | Get all data from this article, Assay Method |
|---|
| |
| Aminopeptidase N |
|---|
| Name: | Aminopeptidase N |
|---|
| Synonyms: | AMPN_PIG | ANPEP | AP-M | AP-N | Alanyl aminopeptidase | Aminopeptidase M | Aminopeptidase N (APN) | CD_antigen=CD13 | Microsomal aminopeptidase | gp130 | pAPN |
|---|
| Type: | Protein |
|---|
| Mol. Mass.: | 108810.25 |
|---|
| Organism: | Sus scrofa (Pig) |
|---|
| Description: | P15145 |
|---|
| Residue: | 963 |
|---|
| Sequence: | MAKGFYISKALGILGILLGVAAVATIIALSVVYAQEKNKNAEHVPQAPTSPTITTTAAIT
LDQSKPWNRYRLPTTLLPDSYNVTLRPYLTPNADGLYIFKGKSIVRLLCQEPTDVIIIHS
KKLNYTTQGHMVVLRGVGDSQVPEIDRTELVELTEYLVVHLKGSLQPGHMYEMESEFQGE
LADDLAGFYRSEYMEGNVKKVLATTQMQSTDARKSFPCFDEPAMKATFNITLIHPNNLTA
LSNMPPKGSSTPLAEDPNWSVTEFETTPVMSTYLLAYIVSEFQSVNETAQNGVLIRIWAR
PNAIAEGHGMYALNVTGPILNFFANHYNTSYPLPKSDQIALPDFNAGAMENWGLVTYREN
ALLFDPQSSSISNKERVVTVIAHELAHQWFGNLVTLAWWNDLWLNEGFASYVEYLGADHA
EPTWNLKDLIVPGDVYRVMAVDALASSHPLTTPAEEVNTPAQISEMFDSISYSKGASVIR
MLSNFLTEDLFKEGLASYLHAFAYQNTTYLDLWEHLQKAVDAQTSIRLPDTVRAIMDRWT
LQMGFPVITVDTKTGNISQKHFLLDSESNVTRSSAFDYLWIVPISSIKNGVMQDHYWLRD
VSQAQNDLFKTASDDWVLLNVNVTGYFQVNYDEDNWRMIQHQLQTNLSVIPVINRAQVIY
DSFNLATAHMVPVTLALDNTLFLNGEKEYMPWQAALSSLSYFSLMFDRSEVYGPMKKYLR
KQVEPLFQHFETLTKNWTERPENLMDQYSEINAISTACSNGLPQCENLAKTLFDQWMSDP
ENNPIHPNLRSTIYCNAIAQGGQDQWDFAWGQLQQAQLVNEADKLRSALACSNEVWLLNR
YLGYTLNPDLIRKQDATSTINSIASNVIGQPLAWDFVQSNWKKLFQDYGGGSFSFSNLIQ
GVTRRFSSEFELQQLEQFKKNNMDVGFGSGTRALEQALEKTKANIKWVKENKEVVLNWFI
EHS
|
|
|
|---|
| BDBM50024594 |
|---|
| n/a |
|---|
| Name | BDBM50024594 |
|---|
| Synonyms: | (R)-1-Ammonium-3-methyl-butane-1-phosphonic acid anion | CHEMBL40422 | LEUCINE PHOSPHONIC ACID |
|---|
| Type | Small organic molecule |
|---|
| Emp. Form. | C5H14NO3P |
|---|
| Mol. Mass. | 167.1433 |
|---|
| SMILES | CC(C)C[C@H]([NH3+])P(O)([O-])=O |
|---|
| Structure | 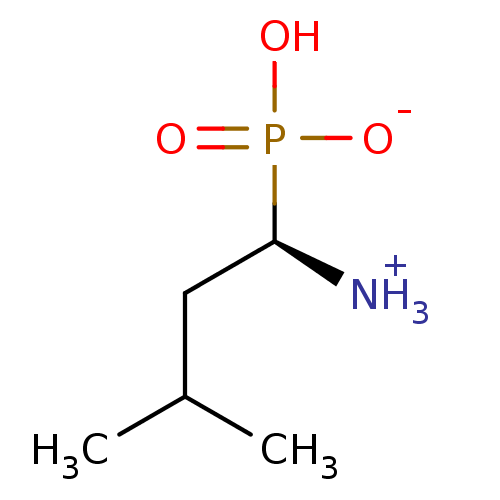
|
|---|


 Search and Browse
Search and Browse
 Download
Download
 Enter Data
Enter Data

 Vassiliou, S; Weglarz-Tomczak, E; Berlicki, L; Pawelczak, M; Nocek, B; Mulligan, R; Joachimiak, A; Mucha, A Structure-guided, single-point modifications in the phosphinic dipeptide structure yield highly potent and selective inhibitors of neutral aminopeptidases. J Med Chem57:8140-51 (2014) [PubMed] Article
Vassiliou, S; Weglarz-Tomczak, E; Berlicki, L; Pawelczak, M; Nocek, B; Mulligan, R; Joachimiak, A; Mucha, A Structure-guided, single-point modifications in the phosphinic dipeptide structure yield highly potent and selective inhibitors of neutral aminopeptidases. J Med Chem57:8140-51 (2014) [PubMed] Article