| Reaction Details |
|---|
 | Report a problem with these data |
| Target | Heat shock protein HSP 90-beta |
|---|
| Ligand | BDBM50165268 |
|---|
| Substrate/Competitor | n/a |
|---|
| Meas. Tech. | ChEMBL_1478162 (CHEMBL3427923) |
|---|
| IC50 | >500000±n/a nM |
|---|
| Citation |  Patel, HJ; Patel, PD; Ochiana, SO; Yan, P; Sun, W; Patel, MR; Shah, SK; Tramentozzi, E; Brooks, J; Bolaender, A; Shrestha, L; Stephani, R; Finotti, P; Leifer, C; Li, Z; Gewirth, DT; Taldone, T; Chiosis, G Structure-activity relationship in a purine-scaffold compound series with selectivity for the endoplasmic reticulum Hsp90 paralog Grp94. J Med Chem58:3922-43 (2015) [PubMed] Article Patel, HJ; Patel, PD; Ochiana, SO; Yan, P; Sun, W; Patel, MR; Shah, SK; Tramentozzi, E; Brooks, J; Bolaender, A; Shrestha, L; Stephani, R; Finotti, P; Leifer, C; Li, Z; Gewirth, DT; Taldone, T; Chiosis, G Structure-activity relationship in a purine-scaffold compound series with selectivity for the endoplasmic reticulum Hsp90 paralog Grp94. J Med Chem58:3922-43 (2015) [PubMed] Article |
|---|
| More Info.: | Get all data from this article, Assay Method |
|---|
| |
| Heat shock protein HSP 90-beta |
|---|
| Name: | Heat shock protein HSP 90-beta |
|---|
| Synonyms: | HS90B_HUMAN | HSP 84 | HSP 90 | HSP90AB1 | HSP90B | HSPC2 | HSPCB | Heat Shock Protein 90 (Hsp90) | Heat shock protein HSP 90 (Hsp90) |
|---|
| Type: | Molecular Chaperone |
|---|
| Mol. Mass.: | 83229.45 |
|---|
| Organism: | Homo sapiens (Human) |
|---|
| Description: | P08238 |
|---|
| Residue: | 724 |
|---|
| Sequence: | MPEEVHHGEEEVETFAFQAEIAQLMSLIINTFYSNKEIFLRELISNASDALDKIRYESLT
DPSKLDSGKELKIDIIPNPQERTLTLVDTGIGMTKADLINNLGTIAKSGTKAFMEALQAG
ADISMIGQFGVGFYSAYLVAEKVVVITKHNDDEQYAWESSAGGSFTVRADHGEPIGRGTK
VILHLKEDQTEYLEERRVKEVVKKHSQFIGYPITLYLEKEREKEISDDEAEEEKGEKEEE
DKDDEEKPKIEDVGSDEEDDSGKDKKKKTKKIKEKYIDQEELNKTKPIWTRNPDDITQEE
YGEFYKSLTNDWEDHLAVKHFSVEGQLEFRALLFIPRRAPFDLFENKKKKNNIKLYVRRV
FIMDSCDELIPEYLNFIRGVVDSEDLPLNISREMLQQSKILKVIRKNIVKKCLELFSELA
EDKENYKKFYEAFSKNLKLGIHEDSTNRRRLSELLRYHTSQSGDEMTSLSEYVSRMKETQ
KSIYYITGESKEQVANSAFVERVRKRGFEVVYMTEPIDEYCVQQLKEFDGKSLVSVTKEG
LELPEDEEEKKKMEESKAKFENLCKLMKEILDKKVEKVTISNRLVSSPCCIVTSTYGWTA
NMERIMKAQALRDNSTMGYMMAKKHLEINPDHPIVETLRQKAEADKNDKAVKDLVVLLFE
TALLSSGFSLEDPQTHSNRIYRMIKLGLGIDEDEVAAEEPNAAVPDEIPPLEGDEDASRM
EEVD
|
|
|
|---|
| BDBM50165268 |
|---|
| n/a |
|---|
| Name | BDBM50165268 |
|---|
| Synonyms: | 8-(3,5-Dichloro-phenylsulfanyl)-9-pent-4-ynyl-9H-purin-6-ylamine | CHEMBL192013 |
|---|
| Type | Small organic molecule |
|---|
| Emp. Form. | C16H13Cl2N5S |
|---|
| Mol. Mass. | 378.279 |
|---|
| SMILES | Nc1ncnc2n(CCCC#C)c(Sc3cc(Cl)cc(Cl)c3)nc12 |
|---|
| Structure | 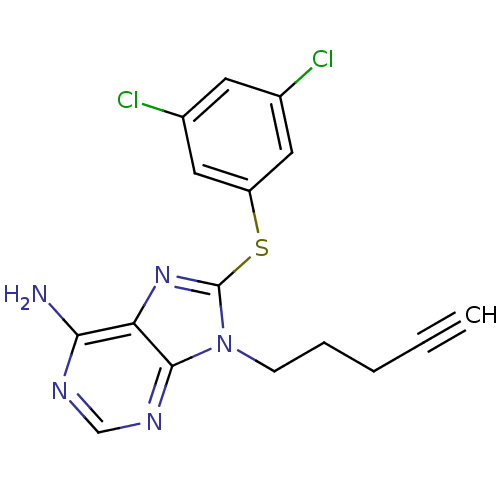
|
|---|


 Search and Browse
Search and Browse
 Download
Download
 Enter Data
Enter Data

 Patel, HJ; Patel, PD; Ochiana, SO; Yan, P; Sun, W; Patel, MR; Shah, SK; Tramentozzi, E; Brooks, J; Bolaender, A; Shrestha, L; Stephani, R; Finotti, P; Leifer, C; Li, Z; Gewirth, DT; Taldone, T; Chiosis, G Structure-activity relationship in a purine-scaffold compound series with selectivity for the endoplasmic reticulum Hsp90 paralog Grp94. J Med Chem58:3922-43 (2015) [PubMed] Article
Patel, HJ; Patel, PD; Ochiana, SO; Yan, P; Sun, W; Patel, MR; Shah, SK; Tramentozzi, E; Brooks, J; Bolaender, A; Shrestha, L; Stephani, R; Finotti, P; Leifer, C; Li, Z; Gewirth, DT; Taldone, T; Chiosis, G Structure-activity relationship in a purine-scaffold compound series with selectivity for the endoplasmic reticulum Hsp90 paralog Grp94. J Med Chem58:3922-43 (2015) [PubMed] Article