| Reaction Details |
|---|
 | Report a problem with these data |
| Target | Polyamine deacetylase HDAC10 |
|---|
| Ligand | BDBM50113971 |
|---|
| Substrate/Competitor | n/a |
|---|
| Meas. Tech. | ChEMBL_1510871 (CHEMBL3606707) |
|---|
| IC50 | >100000±n/a nM |
|---|
| Citation |  Marson, CM; Matthews, CJ; Atkinson, SJ; Lamadema, N; Thomas, NS Potent and Selective Inhibitors of Histone Deacetylase-3 Containing Chiral Oxazoline Capping Groups and a N-(2-Aminophenyl)-benzamide Binding Unit. J Med Chem58:6803-18 (2015) [PubMed] Article Marson, CM; Matthews, CJ; Atkinson, SJ; Lamadema, N; Thomas, NS Potent and Selective Inhibitors of Histone Deacetylase-3 Containing Chiral Oxazoline Capping Groups and a N-(2-Aminophenyl)-benzamide Binding Unit. J Med Chem58:6803-18 (2015) [PubMed] Article |
|---|
| More Info.: | Get all data from this article, Assay Method |
|---|
| |
| Polyamine deacetylase HDAC10 |
|---|
| Name: | Polyamine deacetylase HDAC10 |
|---|
| Synonyms: | HD10 | HDA10_HUMAN | HDAC10 | Histone deacetylase | Histone deacetylase 10 | Human HDAC10 |
|---|
| Type: | Chromatin regulator; hydrolase; repressor |
|---|
| Mol. Mass.: | 71431.89 |
|---|
| Organism: | Homo sapiens (Human) |
|---|
| Description: | Q969S8 |
|---|
| Residue: | 669 |
|---|
| Sequence: | MGTALVYHEDMTATRLLWDDPECEIERPERLTAALDRLRQRGLEQRCLRLSAREASEEEL
GLVHSPEYVSLVRETQVLGKEELQALSGQFDAIYFHPSTFHCARLAAGAGLQLVDAVLTG
AVQNGLALVRPPGHHGQRAAANGFCVFNNVAIAAAHAKQKHGLHRILVVDWDVHHGQGIQ
YLFEDDPSVLYFSWHRYEHGRFWPFLRESDADAVGRGQGLGFTVNLPWNQVGMGNADYVA
AFLHLLLPLAFEFDPELVLVSAGFDSAIGDPEGQMQATPECFAHLTQLLQVLAGGRVCAV
LEGGYHLESLAESVCMTVQTLLGDPAPPLSGPMAPCQSALESIQSARAAQAPHWKSLQQQ
DVTAVPMSPSSHSPEGRPPPLLPGGPVCKAAASAPSSLLDQPCLCPAPSVRTAVALTTPD
ITLVLPPDVIQQEASALREETEAWARPHESLAREEALTALGKLLYLLDGMLDGQVNSGIA
ATPASAAAATLDVAVRRGLSHGAQRLLCVALGQLDRPPDLAHDGRSLWLNIRGKEAAALS
MFHVSTPLPVMTGGFLSCILGLVLPLAYGFQPDLVLVALGPGHGLQGPHAALLAAMLRGL
AGGRVLALLEENSTPQLAGILARVLNGEAPPSLGPSSVASPEDVQALMYLRGQLEPQWKM
LQCHPHLVA
|
|
|
|---|
| BDBM50113971 |
|---|
| n/a |
|---|
| Name | BDBM50113971 |
|---|
| Synonyms: | CHEMBL3605506 |
|---|
| Type | Small organic molecule |
|---|
| Emp. Form. | C29H26N4O2 |
|---|
| Mol. Mass. | 462.5423 |
|---|
| SMILES | Nc1ccccc1NC(=O)c1ccc(CNC2=N[C@@H]([C@@H](O2)c2ccccc2)c2ccccc2)cc1 |r,t:17| |
|---|
| Structure | 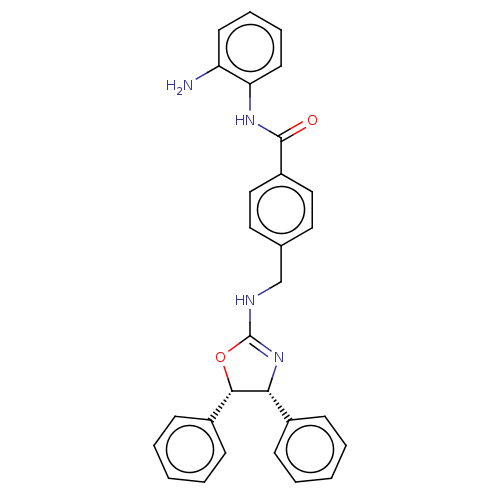
|
|---|


 Search and Browse
Search and Browse
 Download
Download
 Enter Data
Enter Data

 Marson, CM; Matthews, CJ; Atkinson, SJ; Lamadema, N; Thomas, NS Potent and Selective Inhibitors of Histone Deacetylase-3 Containing Chiral Oxazoline Capping Groups and a N-(2-Aminophenyl)-benzamide Binding Unit. J Med Chem58:6803-18 (2015) [PubMed] Article
Marson, CM; Matthews, CJ; Atkinson, SJ; Lamadema, N; Thomas, NS Potent and Selective Inhibitors of Histone Deacetylase-3 Containing Chiral Oxazoline Capping Groups and a N-(2-Aminophenyl)-benzamide Binding Unit. J Med Chem58:6803-18 (2015) [PubMed] Article