| Reaction Details |
|---|
 | Report a problem with these data |
| Target | cAMP-specific 3',5'-cyclic phosphodiesterase 4A/4B/4C/4D |
|---|
| Ligand | BDBM50346088 |
|---|
| Substrate/Competitor | n/a |
|---|
| Meas. Tech. | ChEMBL_157065 (CHEMBL764847) |
|---|
| Ki | 38±n/a nM |
|---|
| Citation |  Napoletano, M; Norcini, G; Pellacini, F; Marchini, F; Morazzoni, G; Fattori, R; Ferlenga, P; Pradella, L Phthalazine PDE4 inhibitors. Part 3: the synthesis and in vitro evaluation of derivatives with a hydrogen bond acceptor. Bioorg Med Chem Lett12:5-8 (2002) [PubMed] Napoletano, M; Norcini, G; Pellacini, F; Marchini, F; Morazzoni, G; Fattori, R; Ferlenga, P; Pradella, L Phthalazine PDE4 inhibitors. Part 3: the synthesis and in vitro evaluation of derivatives with a hydrogen bond acceptor. Bioorg Med Chem Lett12:5-8 (2002) [PubMed] |
|---|
| More Info.: | Get all data from this article, Assay Method |
|---|
| |
| cAMP-specific 3',5'-cyclic phosphodiesterase 4A/4B/4C/4D |
|---|
| Name: | cAMP-specific 3',5'-cyclic phosphodiesterase 4A/4B/4C/4D |
|---|
| Synonyms: | Phosphodiesterase 4 |
|---|
| Type: | n/a |
|---|
| Mol. Mass.: | n/a |
|---|
| Description: | ASSAY_ID of ChEMBL is 788178 |
|---|
| Components: | This complex has 4 components. |
|---|
| Component 1 |
| Name: | cAMP-specific 3',5'-cyclic phosphodiesterase 4A |
|---|
| Synonyms: | 3',5'-cyclic phosphodiesterase | DPDE2 | PDE46 | PDE4A | PDE4A_HUMAN | Phosphodiesterase 4 (PDE4) | Phosphodiesterase 4A | Phosphodiesterase 4A (PDE4) |
|---|
| Type: | Enzyme |
|---|
| Mol. Mass.: | 98113.27 |
|---|
| Organism: | Homo sapiens (Human) |
|---|
| Description: | P27815 |
|---|
| Residue: | 886 |
|---|
| Sequence: | MEPPTVPSERSLSLSLPGPREGQATLKPPPQHLWRQPRTPIRIQQRGYSDSAERAERERQ
PHRPIERADAMDTSDRPGLRTTRMSWPSSFHGTGTGSGGAGGGSSRRFEAENGPTPSPGR
SPLDSQASPGLVLHAGAATSQRRESFLYRSDSDYDMSPKTMSRNSSVTSEAHAEDLIVTP
FAQVLASLRSVRSNFSLLTNVPVPSNKRSPLGGPTPVCKATLSEETCQQLARETLEELDW
CLEQLETMQTYRSVSEMASHKFKRMLNRELTHLSEMSRSGNQVSEYISTTFLDKQNEVEI
PSPTMKEREKQQAPRPRPSQPPPPPVPHLQPMSQITGLKKLMHSNSLNNSNIPRFGVKTD
QEELLAQELENLNKWGLNIFCVSDYAGGRSLTCIMYMIFQERDLLKKFRIPVDTMVTYML
TLEDHYHADVAYHNSLHAADVLQSTHVLLATPALDAVFTDLEILAALFAAAIHDVDHPGV
SNQFLINTNSELALMYNDESVLENHHLAVGFKLLQEDNCDIFQNLSKRQRQSLRKMVIDM
VLATDMSKHMTLLADLKTMVETKKVTSSGVLLLDNYSDRIQVLRNMVHCADLSNPTKPLE
LYRQWTDRIMAEFFQQGDRERERGMEISPMCDKHTASVEKSQVGFIDYIVHPLWETWADL
VHPDAQEILDTLEDNRDWYYSAIRQSPSPPPEEESRGPGHPPLPDKFQFELTLEEEEEEE
ISMAQIPCTAQEALTAQGLSGVEEALDATIAWEASPAQESLEVMAQEASLEAELEAVYLT
QQAQSTGSAPVAPDEFSSREEFVVAVSHSSPSALALQSPLLPAWRTLSVSEHAPGLPGLP
STAAEVEAQREHQAAKRACSACAGTFGEDTSALPAPGGGGSGGDPT
|
|
|
|---|
| Component 2 |
| Name: | cAMP-specific 3',5'-cyclic phosphodiesterase 4D |
|---|
| Synonyms: | 3',5'-cyclic phosphodiesterase | DPDE3 | PDE43 | PDE4D | PDE4D_HUMAN | Phosphodiesterase 4D |
|---|
| Type: | Protein |
|---|
| Mol. Mass.: | 91092.69 |
|---|
| Organism: | Homo sapiens (Human) |
|---|
| Description: | Q08499 |
|---|
| Residue: | 809 |
|---|
| Sequence: | MEAEGSSAPARAGSGEGSDSAGGATLKAPKHLWRHEQHHQYPLRQPQFRLLHPHHHLPPP
PPPSPQPQPQCPLQPPPPPPLPPPPPPPGAARGRYASSGATGRVRHRGYSDTERYLYCRA
MDRTSYAVETGHRPGLKKSRMSWPSSFQGLRRFDVDNGTSAGRSPLDPMTSPGSGLILQA
NFVHSQRRESFLYRSDSDYDLSPKSMSRNSSIASDIHGDDLIVTPFAQVLASLRTVRNNF
AALTNLQDRAPSKRSPMCNQPSINKATITEEAYQKLASETLEELDWCLDQLETLQTRHSV
SEMASNKFKRMLNRELTHLSEMSRSGNQVSEFISNTFLDKQHEVEIPSPTQKEKEKKKRP
MSQISGVKKLMHSSSLTNSSIPRFGVKTEQEDVLAKELEDVNKWGLHVFRIAELSGNRPL
TVIMHTIFQERDLLKTFKIPVDTLITYLMTLEDHYHADVAYHNNIHAADVVQSTHVLLST
PALEAVFTDLEILAAIFASAIHDVDHPGVSNQFLINTNSELALMYNDSSVLENHHLAVGF
KLLQEENCDIFQNLTKKQRQSLRKMVIDIVLATDMSKHMNLLADLKTMVETKKVTSSGVL
LLDNYSDRIQVLQNMVHCADLSNPTKPLQLYRQWTDRIMEEFFRQGDRERERGMEISPMC
DKHNASVEKSQVGFIDYIVHPLWETWADLVHPDAQDILDTLEDNREWYQSTIPQSPSPAP
DDPEEGRQGQTEKFQFELTLEEDGESDTEKDSGSQVEEDTSCSDSKTLCTQDSESTEIPL
DEQVEEEAVGEEEESQPEACVIDDRSPDT
|
|
|
|---|
| Component 3 |
| Name: | cAMP-specific 3',5'-cyclic phosphodiesterase 4B |
|---|
| Synonyms: | 3',5'-cyclic phosphodiesterase | DPDE4 | Isoform PDE4B1 | PDE32 | PDE4B | PDE4B1 | PDE4B_HUMAN | Phosphodiesterase 4B | Phosphodiesterase 4B (PDE4B) | Phosphodiesterase 4B (PDE4B1) | Phosphodiesterase Type 4 (PDE4B) |
|---|
| Type: | Protein |
|---|
| Mol. Mass.: | 83318.87 |
|---|
| Organism: | Homo sapiens (Human) |
|---|
| Description: | Q07343 |
|---|
| Residue: | 736 |
|---|
| Sequence: | MKKSRSVMTVMADDNVKDYFECSLSKSYSSSSNTLGIDLWRGRRCCSGNLQLPPLSQRQS
ERARTPEGDGISRPTTLPLTTLPSIAITTVSQECFDVENGPSPGRSPLDPQASSSAGLVL
HATFPGHSQRRESFLYRSDSDYDLSPKAMSRNSSLPSEQHGDDLIVTPFAQVLASLRSVR
NNFTILTNLHGTSNKRSPAASQPPVSRVNPQEESYQKLAMETLEELDWCLDQLETIQTYR
SVSEMASNKFKRMLNRELTHLSEMSRSGNQVSEYISNTFLDKQNDVEIPSPTQKDREKKK
KQQLMTQISGVKKLMHSSSLNNTSISRFGVNTENEDHLAKELEDLNKWGLNIFNVAGYSH
NRPLTCIMYAIFQERDLLKTFRISSDTFITYMMTLEDHYHSDVAYHNSLHAADVAQSTHV
LLSTPALDAVFTDLEILAAIFAAAIHDVDHPGVSNQFLINTNSELALMYNDESVLENHHL
AVGFKLLQEEHCDIFMNLTKKQRQTLRKMVIDMVLATDMSKHMSLLADLKTMVETKKVTS
SGVLLLDNYTDRIQVLRNMVHCADLSNPTKSLELYRQWTDRIMEEFFQQGDKERERGMEI
SPMCDKHTASVEKSQVGFIDYIVHPLWETWADLVQPDAQDILDTLEDNRNWYQSMIPQSP
SPPLDEQNRDCQGLMEKFQFELTLDEEDSEGPEKEGEGHSYFSSTKTLCVIDPENRDSLG
ETDIDIATEDKSPVDT
|
|
|
|---|
| Component 4 |
| Name: | cAMP-specific 3',5'-cyclic phosphodiesterase 4C |
|---|
| Synonyms: | 3',5'-cyclic phosphodiesterase | DPDE1 | NP_000914.2 | PDE21 | PDE4C | PDE4C1 |
|---|
| Type: | Enzyme Catalytic Domain |
|---|
| Mol. Mass.: | 79875.54 |
|---|
| Organism: | Homo sapiens (Human) |
|---|
| Description: | Q08493 |
|---|
| Residue: | 712 |
|---|
| Sequence: | MENLGVGEGAEACSRLSRSRGRHSMTRAPKHLWRQPRRPIRIQQRFYSDPDKSAGCRERD
LSPRPELRKSRLSWPVSSCRRFDLENGLSCGRRALDPQSSPGLGRIMQAPVPHSQRRESF
LYRSDSDYELSPKAMSRNSSVASDLHGEDMIVTPFAQVLASLRTVRSNVAALARQQCLGA
AKQGPVGNPSSSNQLPPAEDTGQKLALETLDELDWCLDQLETLQTRHSVGEMASNKFKRI
LNRELTHLSETSRSGNQVSEYISRTFLDQQTEVELPKVTAEEAPQPMSRISGLHGLCHSA
SLSSATVPRFGVQTDQEEQLAKELEDTNKWGLDVFKVAELSGNRPLTAIIFSIFQERDLL
KTFQIPADTLATYLLMLEGHYHANVAYHNSLHAADVAQSTHVLLATPALEAVFTDLEILA
ALFASAIHDVDHPGVSNQFLINTNSELALMYNDASVLENHHLAVGFKLLQAENCDIFQNL
SAKQRLSLRRMVIDMVLATDMSKHMNLLADLKTMVETKKVTSLGVLLLDNYSDRIQVLQN
LVHCADLSNPTKPLPLYRQWTDRIMAEFFQQGDRERESGLDISPMCDKHTASVEKSQVGF
IDYIAHPLWETWADLVHPDAQDLLDTLEDNREWYQSKIPRSPSDLTNPERDGPDRFQFEL
TLEEAEEEDEEEEEEGEETALAKEALELPDTELLSPEAGPDPGDLPLDNQRT
|
|
|
|---|
| BDBM50346088 |
|---|
| n/a |
|---|
| Name | BDBM50346088 |
|---|
| Synonyms: | (1r,4r)-4-cyano-4-(3-(cyclopentyloxy)-4-methoxyphenyl)cyclohexanecarboxylic acid | 4-Cyano-3-cyclopentyloxy-4-(4-methoxy-phenyl)-cyclohexanecarboxylic acid | 4-Cyano-4-(3-cyclopentyloxy-4-methoxy-phenyl)-cyclohexanecarboxylic acid | 4-Cyano-4-(3-cyclopentyloxy-4-methoxy-phenyl)-cyclohexanecarboxylic acid (Ariflo) | 4-Cyano-4-(3-cyclopentyloxy-4-methoxy-phenyl)-cyclohexanecarboxylic acid methyl ester | 4-Cyano-4-(3-cyclopentyloxy-4-methoxy-phenyl)-cyclohexanecarboxylic acid(Ariflo) | 6-(3,4-Dimethoxy-phenyl)-5-methyl-4,5-dihydro-2H-pyridazin-3-one | Ariflo | CHEMBL511115 | SB-207499 |
|---|
| Type | Small organic molecule |
|---|
| Emp. Form. | C20H25NO4 |
|---|
| Mol. Mass. | 343.4168 |
|---|
| SMILES | COc1ccc(cc1OC1CCCC1)[C@]1(CC[C@@H](CC1)C(O)=O)C#N |r,wU:17.22,14.25,(4.79,2.97,;3.46,3.74,;2.12,2.97,;.79,3.74,;-.54,2.97,;-.54,1.43,;.79,.66,;2.12,1.43,;3.46,.66,;3.46,-.88,;4.7,-1.79,;4.23,-3.25,;2.69,-3.25,;2.21,-1.79,;-1.88,.66,;-3.42,.71,;-4.23,-.6,;-3.51,-1.96,;-1.97,-2.01,;-1.16,-.7,;-4.33,-3.26,;-3.6,-4.62,;-5.87,-3.21,;-1.77,2.19,;-1.66,3.73,)| |
|---|
| Structure | 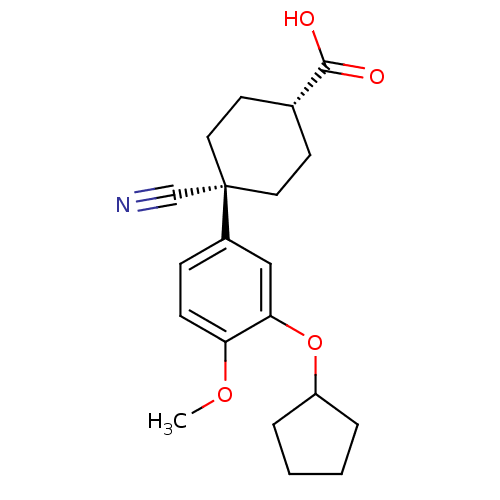
|
|---|


 Search and Browse
Search and Browse
 Download
Download
 Enter Data
Enter Data

 Napoletano, M; Norcini, G; Pellacini, F; Marchini, F; Morazzoni, G; Fattori, R; Ferlenga, P; Pradella, L Phthalazine PDE4 inhibitors. Part 3: the synthesis and in vitro evaluation of derivatives with a hydrogen bond acceptor. Bioorg Med Chem Lett12:5-8 (2002) [PubMed]
Napoletano, M; Norcini, G; Pellacini, F; Marchini, F; Morazzoni, G; Fattori, R; Ferlenga, P; Pradella, L Phthalazine PDE4 inhibitors. Part 3: the synthesis and in vitro evaluation of derivatives with a hydrogen bond acceptor. Bioorg Med Chem Lett12:5-8 (2002) [PubMed]