| Reaction Details |
|---|
 | Report a problem with these data |
| Target | Protein mono-ADP-ribosyltransferase PARP12 |
|---|
| Ligand | BDBM50446130 |
|---|
| Substrate/Competitor | n/a |
|---|
| Meas. Tech. | ChEBML_1646313 |
|---|
| IC50 | 6607±n/a nM |
|---|
| Citation |  Thorsell, AG; Ekblad, T; Karlberg, T; L�w, M; Pinto, AF; Tr�saugues, L; Moche, M; Cohen, MS; Sch�ler, H Structural Basis for Potency and Promiscuity in Poly(ADP-ribose) Polymerase (PARP) and Tankyrase Inhibitors. J Med Chem60:1262-1271 (2017) [PubMed] Article Thorsell, AG; Ekblad, T; Karlberg, T; L�w, M; Pinto, AF; Tr�saugues, L; Moche, M; Cohen, MS; Sch�ler, H Structural Basis for Potency and Promiscuity in Poly(ADP-ribose) Polymerase (PARP) and Tankyrase Inhibitors. J Med Chem60:1262-1271 (2017) [PubMed] Article |
|---|
| More Info.: | Get all data from this article, Assay Method |
|---|
| |
| Protein mono-ADP-ribosyltransferase PARP12 |
|---|
| Name: | Protein mono-ADP-ribosyltransferase PARP12 |
|---|
| Synonyms: | ADP-ribosyltransferase diphtheria toxin-like 12 | ARTD12 | PAR12_HUMAN | PARP-12 | PARP12 | Poly [ADP-ribose] polymerase 12 | ZC3HDC1 | Zinc finger CCCH domain-containing protein 1 |
|---|
| Type: | PROTEIN |
|---|
| Mol. Mass.: | 79092.00 |
|---|
| Organism: | Homo sapiens (Human) |
|---|
| Description: | ChEMBL_107665 |
|---|
| Residue: | 701 |
|---|
| Sequence: | MAQAGVVGEVTQVLCAAGGALELPELRRRLRMGLSADALERLLRQRGRFVVAVRAGGAAA
APERVVLAASPLRLCRAHQGSKPGCVGLCAQLHLCRFMVYGACKFLRAGKNCRNSHSLTT
EHNLSVLRTHGVDHLSYNELCQLLFQNDPWLLPEICQHYNKGDGPHGSCAFQKQCIKLHI
CQYFLQGECKFGTSCKRSHDFSNSENLEKLEKLGMSSDLVSRLPTIYRNAHDIKNKSSAP
SRVPPLFVPQGTSERKDSSGSVSPNTLSQEEGDQICLYHIRKSCSFQDKCHRVHFHLPYR
WQFLDRGKWEDLDNMELIEEAYCNPKIERILCSESASTFHSHCLNFNAMTYGATQARRLS
TASSVTKPPHFILTTDWIWYWSDEFGSWQEYGRQGTVHPVTTVSSSDVEKAYLAYCTPGS
DGQAATLKFQAGKHNYELDFKAFVQKNLVYGTTKKVCRRPKYVSPQDVTTMQTCNTKFPG
PKSIPDYWDSSALPDPGFQKITLSSSSEEYQKVWNLFNRTLPFYFVQKIERVQNLALWEV
YQWQKGQMQKQNGGKAVDERQLFHGTSAIFVDAICQQNFDWRVCGVHGTSYGKGSYFARD
AAYSHHYSKSDTQTHTMFLARVLVGEFVRGNASFVRPPAKEGWSNAFYDSCVNSVSDPSI
FVIFEKHQVYPEYVIQYTTSSKPSVTPSILLALGSLFSSRQ
|
|
|
|---|
| BDBM50446130 |
|---|
| n/a |
|---|
| Name | BDBM50446130 |
|---|
| Synonyms: | AG-014699 | AG-14447 | RUCAPARIB CAMSYLATE | Rucaparib |
|---|
| Type | Small organic molecule |
|---|
| Emp. Form. | C19H18FN3O |
|---|
| Mol. Mass. | 323.3641 |
|---|
| SMILES | CNCc1ccc(cc1)-c1[nH]c2cc(F)cc3C(=O)NCCc1c23 |
|---|
| Structure | 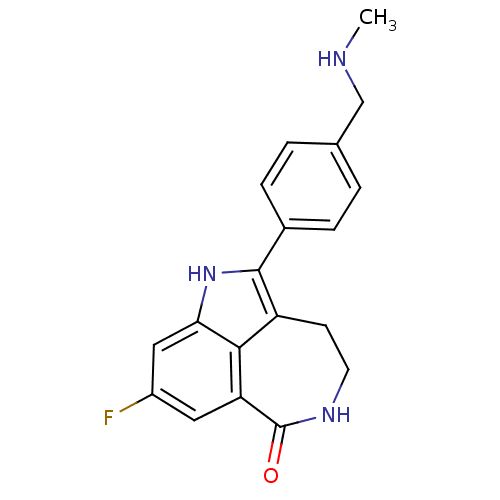
|
|---|


 Search and Browse
Search and Browse
 Download
Download
 Enter Data
Enter Data

 Thorsell, AG; Ekblad, T; Karlberg, T; L�w, M; Pinto, AF; Tr�saugues, L; Moche, M; Cohen, MS; Sch�ler, H Structural Basis for Potency and Promiscuity in Poly(ADP-ribose) Polymerase (PARP) and Tankyrase Inhibitors. J Med Chem60:1262-1271 (2017) [PubMed] Article
Thorsell, AG; Ekblad, T; Karlberg, T; L�w, M; Pinto, AF; Tr�saugues, L; Moche, M; Cohen, MS; Sch�ler, H Structural Basis for Potency and Promiscuity in Poly(ADP-ribose) Polymerase (PARP) and Tankyrase Inhibitors. J Med Chem60:1262-1271 (2017) [PubMed] Article