| Reaction Details |
|---|
 | Report a problem with these data |
| Target | Protein mono-ADP-ribosyltransferase PARP3 |
|---|
| Ligand | BDBM27135 |
|---|
| Substrate/Competitor | n/a |
|---|
| Meas. Tech. | ChEMBL_1646306 (CHEMBL3995362) |
|---|
| IC50 | 2040±n/a nM |
|---|
| Citation |  Thorsell, AG; Ekblad, T; Karlberg, T; L�w, M; Pinto, AF; Tr�saugues, L; Moche, M; Cohen, MS; Sch�ler, H Structural Basis for Potency and Promiscuity in Poly(ADP-ribose) Polymerase (PARP) and Tankyrase Inhibitors. J Med Chem60:1262-1271 (2017) [PubMed] Article Thorsell, AG; Ekblad, T; Karlberg, T; L�w, M; Pinto, AF; Tr�saugues, L; Moche, M; Cohen, MS; Sch�ler, H Structural Basis for Potency and Promiscuity in Poly(ADP-ribose) Polymerase (PARP) and Tankyrase Inhibitors. J Med Chem60:1262-1271 (2017) [PubMed] Article |
|---|
| More Info.: | Get all data from this article, Assay Method |
|---|
| |
| Protein mono-ADP-ribosyltransferase PARP3 |
|---|
| Name: | Protein mono-ADP-ribosyltransferase PARP3 |
|---|
| Synonyms: | ADP-ribosyltransferase diphtheria toxin-like 3 | ADPRT-3 | ADPRT3 | ADPRTL3 | ARTD3 | IRT1 | NAD(+) ADP-ribosyltransferase 3 | PARP-3 | PARP3 | PARP3_HUMAN | Poly [ADP-ribose] polymerase 3 | Poly[ADP-ribose] synthase 3 | hPARP-3 | pADPRT-3 |
|---|
| Type: | PROTEIN |
|---|
| Mol. Mass.: | 60091.40 |
|---|
| Organism: | Homo sapiens (Human) |
|---|
| Description: | ChEMBL_109545 |
|---|
| Residue: | 533 |
|---|
| Sequence: | MAPKPKPWVQTEGPEKKKGRQAGREEDPFRSTAEALKAIPAEKRIIRVDPTCPLSSNPGT
QVYEDYNCTLNQTNIENNNNKFYIIQLLQDSNRFFTCWNRWGRVGEVGQSKINHFTRLED
AKKDFEKKFREKTKNNWAERDHFVSHPGKYTLIEVQAEDEAQEAVVKVDRGPVRTVTKRV
QPCSLDPATQKLITNIFSKEMFKNTMALMDLDVKKMPLGKLSKQQIARGFEALEALEEAL
KGPTDGGQSLEELSSHFYTVIPHNFGHSQPPPINSPELLQAKKDMLLVLADIELAQALQA
VSEQEKTVEEVPHPLDRDYQLLKCQLQLLDSGAPEYKVIQTYLEQTGSNHRCPTLQHIWK
VNQEGEEDRFQAHSKLGNRKLLWHGTNMAVVAAILTSGLRIMPHSGGRVGKGIYFASENS
KSAGYVIGMKCGAHHVGYMFLGEVALGREHHINTDNPSLKSPPPGFDSVIARGHTEPDPT
QDTELELDGQQVVVPQGQPVPCPEFSSSTFSQSEYLIYQESQCRLRYLLEVHL
|
|
|
|---|
| BDBM27135 |
|---|
| n/a |
|---|
| Name | BDBM27135 |
|---|
| Synonyms: | 2-[(2R)-2-methylpyrrolidin-2-yl]-1H-1,3-benzodiazole-4-carboxamide | VELIPARIB | benzimidazole carboxamide, 3a |
|---|
| Type | Small organic molecule |
|---|
| Emp. Form. | C13H16N4O |
|---|
| Mol. Mass. | 244.2923 |
|---|
| SMILES | C[C@@]1(CCCN1)c1nc2cccc(C(N)=O)c2[nH]1 |r| |
|---|
| Structure | 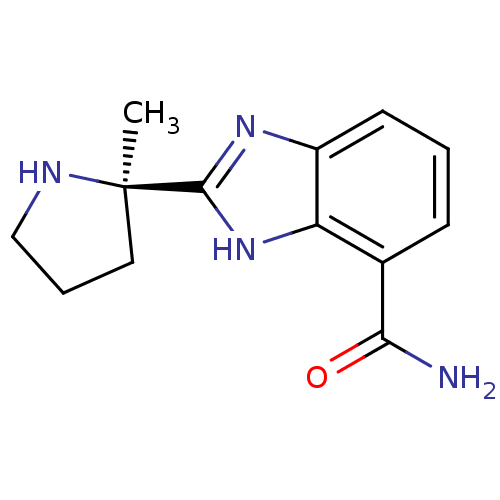
|
|---|


 Search and Browse
Search and Browse
 Download
Download
 Enter Data
Enter Data

 Thorsell, AG; Ekblad, T; Karlberg, T; L�w, M; Pinto, AF; Tr�saugues, L; Moche, M; Cohen, MS; Sch�ler, H Structural Basis for Potency and Promiscuity in Poly(ADP-ribose) Polymerase (PARP) and Tankyrase Inhibitors. J Med Chem60:1262-1271 (2017) [PubMed] Article
Thorsell, AG; Ekblad, T; Karlberg, T; L�w, M; Pinto, AF; Tr�saugues, L; Moche, M; Cohen, MS; Sch�ler, H Structural Basis for Potency and Promiscuity in Poly(ADP-ribose) Polymerase (PARP) and Tankyrase Inhibitors. J Med Chem60:1262-1271 (2017) [PubMed] Article